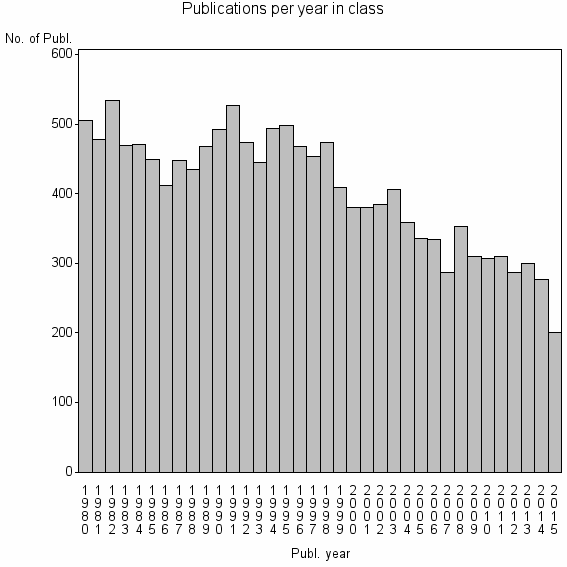
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 501 | 14604 | 39.2 | 67% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 110 | 68452 | JOURNAL OF BACTERIOLOGY//MOLECULAR MICROBIOLOGY//JOURNAL OF MOLECULAR BIOLOGY |
Classes in level below (level 1) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RNA POLYMERASE | Author keyword | 198 | 37% | 3% | 425 |
| 2 | SITE SPECIFIC RECOMBINATION | Author keyword | 91 | 40% | 1% | 181 |
| 3 | H NS | Author keyword | 76 | 62% | 1% | 78 |
| 4 | DNA TRANSPOSITION | Author keyword | 70 | 79% | 0% | 45 |
| 5 | ESCHERICHIA COLI RNA POLYMERASE | Author keyword | 58 | 92% | 0% | 23 |
| 6 | PPGPP | Author keyword | 56 | 57% | 0% | 66 |
| 7 | SERINE RECOMBINASE | Author keyword | 47 | 88% | 0% | 22 |
| 8 | NUSG | Author keyword | 42 | 78% | 0% | 28 |
| 9 | TRP REPRESSOR | Author keyword | 42 | 76% | 0% | 29 |
| 10 | LAC REPRESSOR | Author keyword | 41 | 53% | 0% | 54 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RNA POLYMERASE | 198 | 37% | 3% | 425 | Search RNA+POLYMERASE | Search RNA+POLYMERASE |
| 2 | SITE SPECIFIC RECOMBINATION | 91 | 40% | 1% | 181 | Search SITE+SPECIFIC+RECOMBINATION | Search SITE+SPECIFIC+RECOMBINATION |
| 3 | H NS | 76 | 62% | 1% | 78 | Search H+NS | Search H+NS |
| 4 | DNA TRANSPOSITION | 70 | 79% | 0% | 45 | Search DNA+TRANSPOSITION | Search DNA+TRANSPOSITION |
| 5 | ESCHERICHIA COLI RNA POLYMERASE | 58 | 92% | 0% | 23 | Search ESCHERICHIA+COLI+RNA+POLYMERASE | Search ESCHERICHIA+COLI+RNA+POLYMERASE |
| 6 | PPGPP | 56 | 57% | 0% | 66 | Search PPGPP | Search PPGPP |
| 7 | SERINE RECOMBINASE | 47 | 88% | 0% | 22 | Search SERINE+RECOMBINASE | Search SERINE+RECOMBINASE |
| 8 | NUSG | 42 | 78% | 0% | 28 | Search NUSG | Search NUSG |
| 9 | TRP REPRESSOR | 42 | 76% | 0% | 29 | Search TRP+REPRESSOR | Search TRP+REPRESSOR |
| 10 | LAC REPRESSOR | 41 | 53% | 0% | 54 | Search LAC+REPRESSOR | Search LAC+REPRESSOR |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHAGE MU | 277 | 92% | 1% | 108 |
| 2 | BACTERIOPHAGE MU | 261 | 70% | 1% | 217 |
| 3 | LAC UV5 PROMOTER | 234 | 82% | 1% | 135 |
| 4 | COLI RNA POLYMERASE | 219 | 45% | 2% | 363 |
| 5 | TRIGGER LOOP | 184 | 88% | 1% | 87 |
| 6 | BACTERIOPHAGE LAMBDA | 166 | 30% | 3% | 469 |
| 7 | GAMMA DELTA RESOLVASE | 163 | 80% | 1% | 101 |
| 8 | TRANSCRIPTION INITIATION | 152 | 35% | 2% | 348 |
| 9 | GUANOSINE TETRAPHOSPHATE | 149 | 66% | 1% | 138 |
| 10 | OPEN COMPLEX FORMATION | 149 | 71% | 1% | 120 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOBILE DNA | 1 | 10% | 0% | 12 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| USE OF T7 RNA-POLYMERASE TO DIRECT EXPRESSION OF CLONED GENES | 1990 | 6055 | 16 | 56% |
| (p)ppGpp: Still Magical? | 2008 | 353 | 79 | 80% |
| Transcription activation by catabolite activator protein (CAP) | 1999 | 460 | 98 | 92% |
| Bacterial nucleoid-associated proteins, nucleoid structure and gene expression | 2010 | 195 | 127 | 58% |
| Insertion sequences | 1998 | 854 | 337 | 53% |
| The RpoS-Mediated General Stress Response in Escherichia coli | 2011 | 141 | 161 | 55% |
| Mechanism of Bacterial Transcription Initiation: RNA Polymerase - Promoter Binding, Isomerization to Initiation-Competent Open Complexes, and Initiation of RNA Synthesis | 2011 | 73 | 134 | 93% |
| ppGpp: a global regulator in Escherichia coli | 2005 | 260 | 53 | 89% |
| Mechanisms of site-specific recombination | 2006 | 259 | 136 | 79% |
| Bacterial Transcription Terminators: The RNA 3 '-End Chronicles | 2011 | 66 | 121 | 91% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHYSIOL BACTERIENNE | 22 | 81% | 0.1% | 13 |
| 2 | TRANSCRIPT BIOL | 21 | 75% | 0.1% | 15 |
| 3 | SKIRBALL PROGRAM MOL PATHOGENESIS | 19 | 74% | 0.1% | 14 |
| 4 | SECT MICROBIAL GENET | 15 | 82% | 0.1% | 9 |
| 5 | WAKSMAN | 14 | 14% | 0.6% | 94 |
| 6 | LEIDEN CHEM CELL OBSERV | 14 | 100% | 0.0% | 7 |
| 7 | GENE EXP S REGULAT SECT | 13 | 71% | 0.1% | 10 |
| 8 | MICROBIOL GENET MOL | 12 | 16% | 0.5% | 68 |
| 9 | MORSE MOL GENET | 11 | 43% | 0.1% | 19 |
| 10 | UNITE PHYSICOCHIM MACROMOL BIOL | 9 | 45% | 0.1% | 15 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000018431 | FTSZ//DNAA//CHLOROPLAST DIVISION |
| 2 | 0.0000015746 | NIFA PROTEIN//GLNK//SIGMA54 |
| 3 | 0.0000013908 | RESTRICTION ENDONUCLEASE//RESTRICTION MODIFICATION//RESTRICTION MODIFICATION SYSTEM |
| 4 | 0.0000013268 | Z DNA//JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS//B Z TRANSITION |
| 5 | 0.0000013165 | PHOSPHOTRANSFERASE SYSTEM//LACTOSE PERMEASE//BIOL BAKTERIENPHYSIOL |
| 6 | 0.0000011838 | BACTERIOPHAGE//PHAGE THERAPY//DNA PACKAGING |
| 7 | 0.0000011015 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
| 8 | 0.0000009738 | RIBOSOME//TRANS TRANSLATION//TMRNA |
| 9 | 0.0000008338 | ENTEROCOCCUS//ENTEROCOCCI//VANA |
| 10 | 0.0000008231 | RNASE P//AMINOACYL TRNA SYNTHETASE//TRNA |