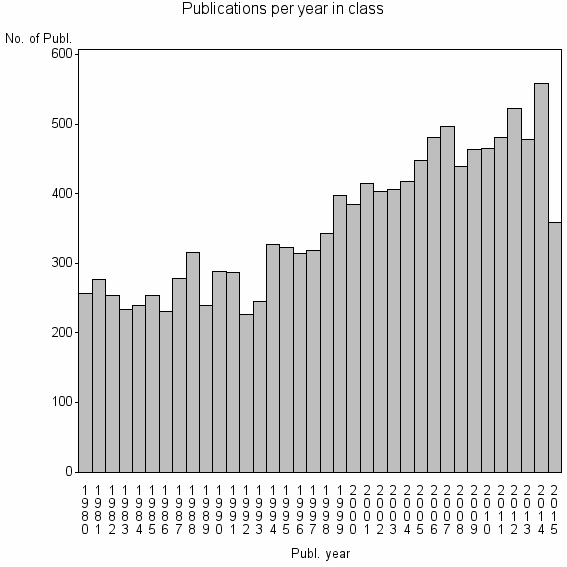
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 646 | 12857 | 41.9 | 75% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 136 | 62081 | RNA-A PUBLICATION OF THE RNA SOCIETY//RIBOZYME//NUCLEIC ACIDS RESEARCH |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 1184 | 2699 | MRNA ANALOG//80S RIBOSOME//HUMAN RIBOSOME |
| 2527 | 2146 | ELONGATION FACTOR TU//EF G//ELONGATION FACTOR G |
| 3151 | 1972 | TRANSLATIONAL SELECTION//CODON USAGE//SYNONYMOUS CODON USAGE |
| 5238 | 1584 | ATP DEPENDENT PROTEASES//CLPB//FTSH |
| 6844 | 1360 | SUP35//TRANSLATION TERMINATION//URE3 |
| 12225 | 858 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 14068 | 729 | SD SEQUENCE//RIBOSOMAL PROTEIN S1//SHINE DALGARNO SEQUENCE |
| 14369 | 708 | TRIGGER FACTOR//NASCENT CHAIN//COTRANSLATIONAL FOLDING |
| 18592 | 479 | TRANS TRANSLATION//TMRNA//SMPB |
| 22524 | 322 | OBG//DRG2//BACTERIAL GTPASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RIBOSOME | Author keyword | 234 | 36% | 4% | 528 |
| 2 | TRANS TRANSLATION | Author keyword | 190 | 93% | 1% | 70 |
| 3 | TMRNA | Author keyword | 176 | 85% | 1% | 92 |
| 4 | TRANSLATION TERMINATION | Author keyword | 170 | 73% | 1% | 130 |
| 5 | ATP DEPENDENT PROTEASES | Author keyword | 167 | 85% | 1% | 89 |
| 6 | SUP35 | Author keyword | 131 | 87% | 1% | 65 |
| 7 | CODON USAGE | Author keyword | 107 | 38% | 2% | 228 |
| 8 | SYNONYMOUS CODON USAGE | Author keyword | 93 | 81% | 0% | 56 |
| 9 | TRANSLATIONAL SELECTION | Author keyword | 93 | 87% | 0% | 46 |
| 10 | ELONGATION FACTOR TU | Author keyword | 88 | 68% | 1% | 77 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RIBOSOME | 234 | 36% | 4% | 528 | Search RIBOSOME | Search RIBOSOME |
| 2 | TRANS TRANSLATION | 190 | 93% | 1% | 70 | Search TRANS+TRANSLATION | Search TRANS+TRANSLATION |
| 3 | TMRNA | 176 | 85% | 1% | 92 | Search TMRNA | Search TMRNA |
| 4 | TRANSLATION TERMINATION | 170 | 73% | 1% | 130 | Search TRANSLATION+TERMINATION | Search TRANSLATION+TERMINATION |
| 5 | ATP DEPENDENT PROTEASES | 167 | 85% | 1% | 89 | Search ATP+DEPENDENT+PROTEASES | Search ATP+DEPENDENT+PROTEASES |
| 6 | SUP35 | 131 | 87% | 1% | 65 | Search SUP35 | Search SUP35 |
| 7 | CODON USAGE | 107 | 38% | 2% | 228 | Search CODON+USAGE | Search CODON+USAGE |
| 8 | SYNONYMOUS CODON USAGE | 93 | 81% | 0% | 56 | Search SYNONYMOUS+CODON+USAGE | Search SYNONYMOUS+CODON+USAGE |
| 9 | TRANSLATIONAL SELECTION | 93 | 87% | 0% | 46 | Search TRANSLATIONAL+SELECTION | Search TRANSLATIONAL+SELECTION |
| 10 | ELONGATION FACTOR TU | 88 | 68% | 1% | 77 | Search ELONGATION+FACTOR+TU | Search ELONGATION+FACTOR+TU |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PSI PRION | 638 | 95% | 2% | 212 |
| 2 | ATP DEPENDENT PROTEASES | 391 | 64% | 3% | 388 |
| 3 | DE NOVO APPEARANCE | 385 | 97% | 1% | 107 |
| 4 | AMINOACYL TRANSFER RNA | 354 | 55% | 3% | 442 |
| 5 | ESCHERICHIA COLI RIBOSOMES | 345 | 63% | 3% | 349 |
| 6 | TRANSLATION TERMINATION | 337 | 73% | 2% | 256 |
| 7 | 70S RIBOSOME | 307 | 72% | 2% | 244 |
| 8 | TRANSLATIONAL SELECTION | 304 | 84% | 1% | 168 |
| 9 | ELONGATION FACTOR G | 298 | 64% | 2% | 291 |
| 10 | P SITE | 286 | 81% | 1% | 173 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PRION | 13 | 17% | 1% | 67 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Synonymous but not the same: the causes and consequences of codon bias | 2011 | 253 | 131 | 72% |
| What recent ribosome structures have revealed about the mechanism of translation | 2009 | 254 | 126 | 96% |
| AAA+ Proteases: ATP-Fueled Machines of Protein Destruction | 2011 | 137 | 160 | 74% |
| Ribosome rescue systems in bacteria | 2015 | 2 | 144 | 84% |
| ClpXP, an ATP-powered unfolding and protein-degradation machine | 2012 | 61 | 108 | 92% |
| AAA+ proteins: Have engine, will work | 2005 | 491 | 103 | 56% |
| The tmRNA system for translational surveillance and ribosome rescue | 2007 | 151 | 131 | 91% |
| Ribosome structure and the mechanism of translation | 2002 | 402 | 96 | 86% |
| Selection on Codon Bias | 2008 | 216 | 45 | 64% |
| A structural understanding of the dynamic ribosome machine | 2008 | 153 | 48 | 88% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AG RIBOSOMEN | 63 | 76% | 0.3% | 44 |
| 2 | MOL BIOL RNA | 30 | 35% | 0.5% | 68 |
| 3 | ORG EVOLUC GENOMA | 26 | 61% | 0.2% | 28 |
| 4 | JW WILSON | 17 | 63% | 0.1% | 17 |
| 5 | GENET MOL CHAMPIGNONS | 16 | 56% | 0.2% | 20 |
| 6 | ABT VINGRON | 13 | 80% | 0.1% | 8 |
| 7 | ARRHENIUS S E5 | 13 | 71% | 0.1% | 10 |
| 8 | KENT FUNGAL GRP | 12 | 58% | 0.1% | 14 |
| 9 | HLTH INC | 10 | 50% | 0.1% | 15 |
| 10 | BIOL MCA | 10 | 34% | 0.2% | 25 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000018365 | RNASE P//AMINOACYL TRNA SYNTHETASE//TRNA |
| 2 | 0.0000014802 | IRES//TRANSLATION INITIATION//EIF4E |
| 3 | 0.0000012993 | RIBOZYME//GROUP I INTRON//RIBOSWITCH |
| 4 | 0.0000012176 | NUCLEOLUS//RIBOSOME BIOGENESIS//NUCLEOLAR ORGANIZER REGIONS |
| 5 | 0.0000011817 | DANISH ARCHAEA//METHANOSARCINA BARKERI//ARCHAEA |
| 6 | 0.0000009738 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
| 7 | 0.0000009558 | PROTEIN TRANSLOCATION//PROTEIN IMPORT//SECA |
| 8 | 0.0000006763 | PREDICTION WITH EXPERT ADVICE//UNIVERSAL CODING//GRAPHICAL REPRESENTATION |
| 9 | 0.0000006757 | PHOSPHOLIPIDOSIS//2 DEOXYSTREPTAMINE//AMINOGLYCOSIDES |
| 10 | 0.0000006454 | CELL FREE PROTEIN SYNTHESIS//MEMBRANE PROTEIN//RIBOSOME DISPLAY |