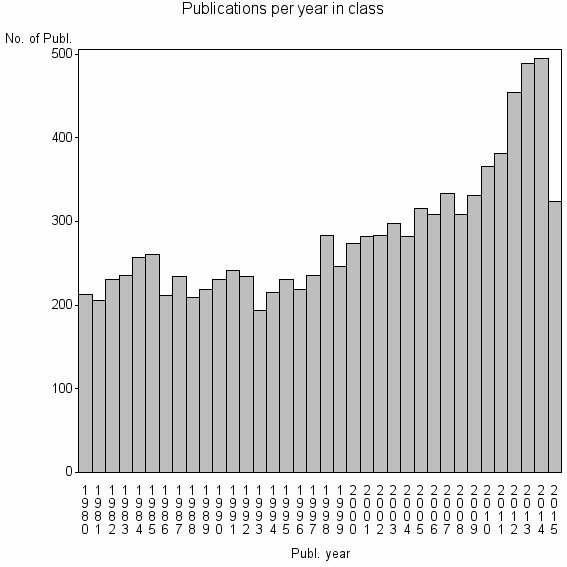
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 979 | 10123 | 42.7 | 73% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 110 | 68452 | JOURNAL OF BACTERIOLOGY//MOLECULAR MICROBIOLOGY//JOURNAL OF MOLECULAR BIOLOGY |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 2855 | 2057 | UMR8041//USM0502//MUREIN |
| 4574 | 1690 | FTSZ//CHLOROPLAST DIVISION//PLASTID DIVISION |
| 6967 | 1346 | DNAA PROTEIN//DNAA//ORIC |
| 8033 | 1225 | BACTERIAL CONJUGATION//COUPLING PROTEIN//R391 |
| 8072 | 1221 | REPLICATION CONTROL//PLASMID R6K//PLASMID RK2 |
| 10581 | 985 | TOXIN ANTITOXIN//TOXIN ANTITOXIN SYSTEM//TOXIN ANTITOXIN SYSTEMS |
| 10618 | 981 | PARB//DNA SEGREGATION//FTSK |
| 15874 | 618 | CAULOBACTER//CTRA//CAULOBACTER CRESCENTUS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | FTSZ | Author keyword | 325 | 77% | 2% | 222 |
| 2 | DNAA | Author keyword | 125 | 79% | 1% | 81 |
| 3 | CHLOROPLAST DIVISION | Author keyword | 109 | 83% | 1% | 62 |
| 4 | ORIC | Author keyword | 105 | 80% | 1% | 66 |
| 5 | DNAA PROTEIN | Author keyword | 103 | 93% | 0% | 39 |
| 6 | PLASTID DIVISION | Author keyword | 97 | 82% | 1% | 56 |
| 7 | BACTERIAL CELL DIVISION | Author keyword | 92 | 84% | 1% | 51 |
| 8 | PLASMID | Journal | 74 | 18% | 4% | 364 |
| 9 | BACTERIAL CONJUGATION | Author keyword | 69 | 71% | 1% | 55 |
| 10 | BACTERIAL CELL BIOL | Address | 69 | 48% | 1% | 105 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | FTSZ | 325 | 77% | 2% | 222 | Search FTSZ | Search FTSZ |
| 2 | DNAA | 125 | 79% | 1% | 81 | Search DNAA | Search DNAA |
| 3 | CHLOROPLAST DIVISION | 109 | 83% | 1% | 62 | Search CHLOROPLAST+DIVISION | Search CHLOROPLAST+DIVISION |
| 4 | ORIC | 105 | 80% | 1% | 66 | Search ORIC | Search ORIC |
| 5 | DNAA PROTEIN | 103 | 93% | 0% | 39 | Search DNAA+PROTEIN | Search DNAA+PROTEIN |
| 6 | PLASTID DIVISION | 97 | 82% | 1% | 56 | Search PLASTID+DIVISION | Search PLASTID+DIVISION |
| 7 | BACTERIAL CELL DIVISION | 92 | 84% | 1% | 51 | Search BACTERIAL+CELL+DIVISION | Search BACTERIAL+CELL+DIVISION |
| 8 | BACTERIAL CONJUGATION | 69 | 71% | 1% | 55 | Search BACTERIAL+CONJUGATION | Search BACTERIAL+CONJUGATION |
| 9 | TOXIN ANTITOXIN | 58 | 74% | 0% | 43 | Search TOXIN+ANTITOXIN | Search TOXIN+ANTITOXIN |
| 10 | TOXIN ANTITOXIN SYSTEM | 49 | 79% | 0% | 31 | Search TOXIN+ANTITOXIN+SYSTEM | Search TOXIN+ANTITOXIN+SYSTEM |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BACTERIAL CELL DIVISION | 533 | 85% | 3% | 280 |
| 2 | Z RING | 501 | 90% | 2% | 217 |
| 3 | F PLASMID | 452 | 86% | 2% | 229 |
| 4 | PROTEIN FTSZ | 376 | 87% | 2% | 181 |
| 5 | TO POLE OSCILLATION | 320 | 90% | 1% | 138 |
| 6 | PROPER PLACEMENT | 313 | 97% | 1% | 91 |
| 7 | RAPID POLE | 307 | 92% | 1% | 123 |
| 8 | INHIBITOR MINC | 302 | 98% | 1% | 81 |
| 9 | CELL DIVISION PROTEIN | 278 | 89% | 1% | 127 |
| 10 | SEPTAL RING | 258 | 89% | 1% | 118 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PLASMID | 74 | 18% | 4% | 364 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| From the regulation of peptidoglycan synthesis to bacterial growth and morphology | 2012 | 156 | 134 | 80% |
| Molecular Mechanisms Underlying Bacterial Persisters | 2014 | 40 | 91 | 77% |
| Bacterial cell division: assembly, maintenance and disassembly of the Z ring | 2009 | 290 | 155 | 93% |
| Persister Cells | 2010 | 357 | 79 | 52% |
| Toxin-Antitoxin Systems as Multilevel Interaction Systems | 2014 | 22 | 124 | 94% |
| Toxin-Antitoxin Systems in Bacteria and Archaea | 2011 | 129 | 109 | 93% |
| FtsZ in Bacterial Cytokinesis: Cytoskeleton and Force Generator All in One | 2010 | 185 | 200 | 86% |
| Assembly dynamics of the bacterial MinCDE system and spatial regulation of the Z ring | 2007 | 233 | 150 | 97% |
| Peptidoglycan structure and architecture | 2008 | 351 | 107 | 66% |
| Bacterial Persistence and Toxin-Antitoxin Loci | 2012 | 91 | 108 | 80% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BACTERIAL CELL BIOL | 69 | 48% | 1.0% | 105 |
| 2 | UMR8041 | 26 | 100% | 0.1% | 11 |
| 3 | USM0502 | 26 | 87% | 0.1% | 13 |
| 4 | ANTIMICROBIAL DISCOVERY | 22 | 58% | 0.2% | 25 |
| 5 | PLATEFORME SPE OMETRIE MASSE PROTE MUSEUM | 17 | 100% | 0.1% | 8 |
| 6 | GENET PROCARYOTES | 11 | 49% | 0.2% | 17 |
| 7 | BIOL ESTRUCT MOL | 8 | 100% | 0.0% | 5 |
| 8 | RECH DEV ERS MOL | 8 | 100% | 0.0% | 5 |
| 9 | RECH DEV ERSITE MOL | 8 | 100% | 0.0% | 5 |
| 10 | LEHRBEREICH MIKROBIELLE GENET | 8 | 62% | 0.1% | 8 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000018431 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
| 2 | 0.0000016133 | ENTEROCOCCUS//ENTEROCOCCI//VANA |
| 3 | 0.0000013657 | MYXOCOCCUS XANTHUS//BDELLOVIBRIO//BDELLOVIBRIO BACTERIOVORUS |
| 4 | 0.0000011742 | CROWN GALL//PHTHORIMAEA OPERCULELLA//AGROBACTERIUM VITIS |
| 5 | 0.0000009778 | L FORMS//OPTICAL BEAM CONDENSER//ABSORPTION DATA |
| 6 | 0.0000009242 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
| 7 | 0.0000008579 | RESTRICTION ENDONUCLEASE//RESTRICTION MODIFICATION//RESTRICTION MODIFICATION SYSTEM |
| 8 | 0.0000008372 | PEPTIDOGLYCAN MONOMER//TEICHOIC ACIDS//TEICHOIC ACID |
| 9 | 0.0000008294 | AUTOTRANSPORTER//PORIN//BACTERIAL CHEMOTAXIS |
| 10 | 0.0000007117 | DISSOLVED DNA//NATURAL TRANSFORMATION//EDNA |