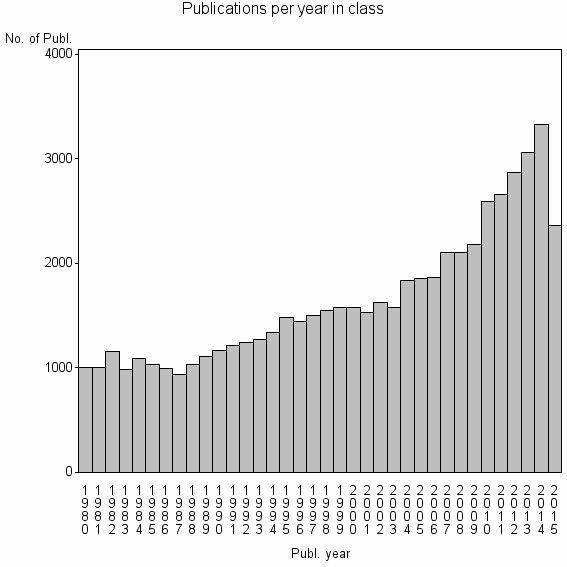
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 151 | 59191 | 46.0 | 78% |
Classes in level above (level 4) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 0 | 3633673 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//MED |
Classes in level below (level 2) |
| ID, lev. below |
Publications | Label for level below |
|---|---|---|
| 181 | 21089 | CHROMATIN//EZH2//NUCLEOSOME |
| 695 | 12311 | CHROMOSOMA//SYNAPTONEMAL COMPLEX//MEIOSIS |
| 1031 | 9720 | NUCLEOLUS//RIBOSOME BIOGENESIS//NUCLEOLAR ORGANIZER REGIONS |
| 1810 | 5735 | NUCLEAR ENVELOPE//NUCLEAR PORE COMPLEX//NUCLEAR LAMINA |
| 2134 | 4592 | GEMININ//DNA REPLICATION//CDC6 |
| 2251 | 4244 | NUCLEAR MATRIX//PROT SECT//MATRIX ATTACHMENT REGION |
| 3610 | 894 | MICROBIAL BIOTECHNOL CELL BIOL//INTERPHASE CHROMATIN//BARLEY CHROMOSOME |
| 3827 | 606 | HUMAN HYALINE CARTILAGE//JITTER FREE//BLUE CHROMOPHORES |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CHROMATIN | Author keyword | 780 | 36% | 3% | 1742 |
| 2 | CHROMOSOMA | Journal | 597 | 42% | 2% | 1091 |
| 3 | NUCLEAR ENVELOPE | Author keyword | 477 | 62% | 1% | 487 |
| 4 | NUCLEAR PORE COMPLEX | Author keyword | 414 | 70% | 1% | 346 |
| 5 | NUCLEAR LAMINA | Author keyword | 358 | 86% | 0% | 179 |
| 6 | NUCLEOSOME | Author keyword | 354 | 58% | 1% | 409 |
| 7 | NUCLEOLUS | Author keyword | 326 | 43% | 1% | 586 |
| 8 | EZH2 | Author keyword | 281 | 65% | 0% | 267 |
| 9 | NUCLEAR MATRIX | Author keyword | 249 | 49% | 1% | 371 |
| 10 | RNA POLYMERASE II | Author keyword | 248 | 48% | 1% | 384 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CHROMATIN | 780 | 36% | 3% | 1742 | Search CHROMATIN | Search CHROMATIN |
| 2 | NUCLEAR ENVELOPE | 477 | 62% | 1% | 487 | Search NUCLEAR+ENVELOPE | Search NUCLEAR+ENVELOPE |
| 3 | NUCLEAR PORE COMPLEX | 414 | 70% | 1% | 346 | Search NUCLEAR+PORE+COMPLEX | Search NUCLEAR+PORE+COMPLEX |
| 4 | NUCLEAR LAMINA | 358 | 86% | 0% | 179 | Search NUCLEAR+LAMINA | Search NUCLEAR+LAMINA |
| 5 | NUCLEOSOME | 354 | 58% | 1% | 409 | Search NUCLEOSOME | Search NUCLEOSOME |
| 6 | NUCLEOLUS | 326 | 43% | 1% | 586 | Search NUCLEOLUS | Search NUCLEOLUS |
| 7 | EZH2 | 281 | 65% | 0% | 267 | Search EZH2 | Search EZH2 |
| 8 | NUCLEAR MATRIX | 249 | 49% | 1% | 371 | Search NUCLEAR+MATRIX | Search NUCLEAR+MATRIX |
| 9 | RNA POLYMERASE II | 248 | 48% | 1% | 384 | Search RNA+POLYMERASE+II | Search RNA+POLYMERASE+II |
| 10 | EMERIN | 236 | 83% | 0% | 131 | Search EMERIN | Search EMERIN |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CHROMATIN | 1703 | 35% | 7% | 3899 |
| 2 | RNA POLYMERASE II | 1557 | 36% | 6% | 3500 |
| 3 | DREIFUSS MUSCULAR DYSTROPHY | 896 | 80% | 1% | 555 |
| 4 | POSITION EFFECT VARIEGATION | 672 | 63% | 1% | 671 |
| 5 | PRE RIBOSOMAL RNA | 657 | 73% | 1% | 502 |
| 6 | CHROMATIN STRUCTURE | 608 | 35% | 2% | 1402 |
| 7 | A TYPE LAMINS | 552 | 86% | 0% | 283 |
| 8 | ORIGIN RECOGNITION COMPLEX | 534 | 63% | 1% | 544 |
| 9 | NUCLEOCYTOPLASMIC TRANSPORT | 525 | 52% | 1% | 722 |
| 10 | PORE COMPLEX | 494 | 54% | 1% | 629 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CHROMOSOMA | 597 | 42% | 2% | 1091 |
| 2 | MOLECULAR AND CELLULAR BIOLOGY | 238 | 10% | 4% | 2211 |
| 3 | GENETICS | 143 | 11% | 2% | 1286 |
| 4 | MOLECULAR CELL | 136 | 16% | 1% | 787 |
| 5 | GENES & DEVELOPMENT | 132 | 13% | 2% | 938 |
| 6 | NUCLEUS-AUSTIN | 132 | 63% | 0% | 134 |
| 7 | NUCLEUS | 92 | 64% | 0% | 89 |
| 8 | CHROMOSOME RESEARCH | 56 | 18% | 0% | 283 |
| 9 | EPIGENETICS & CHROMATIN | 53 | 44% | 0% | 90 |
| 10 | NATURE STRUCTURAL & MOLECULAR BIOLOGY | 37 | 13% | 0% | 262 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Chromatin modifications and their function | 2007 | 3808 | 89 | 63% |
| Translating the histone code | 2001 | 4788 | 80 | 70% |
| The role of chromatin during transcription | 2007 | 1530 | 125 | 93% |
| The Polycomb complex PRC2 and its mark in life | 2011 | 598 | 96 | 81% |
| The complex language of chromatin regulation during transcription | 2007 | 1163 | 48 | 85% |
| Regulation of chromatin by histone modifications | 2011 | 589 | 121 | 63% |
| The Biology of Chromatin Remodeling Complexes | 2009 | 630 | 179 | 83% |
| Histone exchange, chromatin structure and the regulation of transcription | 2015 | 7 | 155 | 94% |
| Regulation of nucleosome dynamics by histone modifications | 2013 | 159 | 101 | 83% |
| Cancer Genome Landscapes | 2013 | 802 | 139 | 17% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CONTROL GENET PROC | 202 | 98% | 0.1% | 51 |
| 2 | PROT SECT | 148 | 93% | 0.1% | 55 |
| 3 | BIOL MOL EUCARYOTE | 126 | 61% | 0.2% | 132 |
| 4 | MOL BIOL CELL 2 | 79 | 73% | 0.1% | 61 |
| 5 | GENE BIOL | 73 | 27% | 0.4% | 229 |
| 6 | NUCLE ACIDS ENZYMOL | 55 | 92% | 0.0% | 22 |
| 7 | RECEPTOR BIOL GENE EXP S | 53 | 38% | 0.2% | 111 |
| 8 | ADOLF BUTENANDT | 38 | 24% | 0.2% | 135 |
| 9 | STRUCT GENOM CONSORTIUM | 37 | 22% | 0.3% | 150 |
| 10 | CYTOL GENET | 35 | 11% | 0.5% | 317 |
Related classes at same level (level 3) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000004567 | TELOMERASE//TELOMERE//FANCONI ANEMIA |
| 2 | 0.0000004149 | RNA-A PUBLICATION OF THE RNA SOCIETY//RIBOZYME//NUCLEIC ACIDS RESEARCH |
| 3 | 0.0000003303 | KINETOCHORE//MITOSIS//KINESIN |
| 4 | 0.0000002940 | TRANSCRIPTION FACTOR//ETS 1//C JUN |
| 5 | 0.0000002834 | DNA METHYLATION//FRAGILE X SYNDROME//RETT SYNDROME |
| 6 | 0.0000002669 | GENOME EDITING//CAS9//CRISPR |
| 7 | 0.0000002586 | MERKEL CELL CARCINOMA//P53//JC VIRUS |
| 8 | 0.0000002410 | BIOINFORMATICS//BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY |
| 9 | 0.0000001943 | CYTOMETRY//ANALYTICAL AND QUANTITATIVE CYTOLOGY AND HISTOLOGY//CYTOMETRY PART A |
| 10 | 0.0000001846 | SACCHAROMYCES CEREVISIAE//XYLANASE//YEAST |