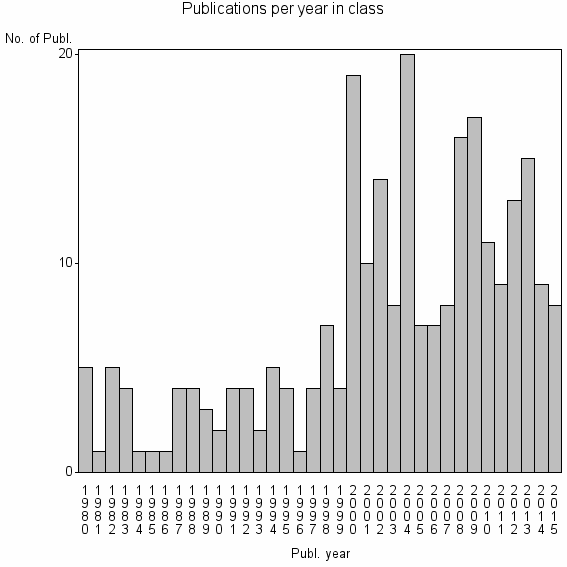
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 24600 | 257 | 33.7 | 70% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1852 | 5569 | LESCH NYHAN SYNDROME//TIAZOFURIN//LESCH NYHAN DISEASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ODCASE | Author keyword | 26 | 100% | 4% | 11 |
| 2 | OROTIDINE 5 MONOPHOSPHATE DECARBOXYLASE | Author keyword | 11 | 69% | 4% | 9 |
| 3 | MOL DESIGN PREFORMULAT | Address | 5 | 42% | 4% | 10 |
| 4 | WEAK LINKAGE | Author keyword | 5 | 63% | 2% | 5 |
| 5 | OROTIDINE MONOPHOSPHATE DECARBOXYLASE | Author keyword | 3 | 100% | 1% | 3 |
| 6 | OMP DECARBOXYLASE | Author keyword | 2 | 50% | 1% | 3 |
| 7 | OROTIDINE | Author keyword | 2 | 40% | 2% | 4 |
| 8 | OROTIDINE 5 MONOPHOSPHATE DECARBOXYLASE | Author keyword | 2 | 43% | 1% | 3 |
| 9 | DE NOVO NUCLEOTIDE SYNTHESIS | Author keyword | 1 | 100% | 1% | 2 |
| 10 | MCLAUGHLIN ROTMAN UHN | Address | 1 | 100% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | OMP DECARBOXYLASE | 206 | 98% | 20% | 52 |
| 2 | PROFICIENT ENZYME | 165 | 82% | 37% | 96 |
| 3 | 5 PHOSPHATE DECARBOXYLASE | 150 | 96% | 18% | 47 |
| 4 | MONOPHOSPHATE DECARBOXYLASE | 137 | 93% | 20% | 52 |
| 5 | 5 MONOPHOSPHATE DECARBOXYLASE | 79 | 94% | 11% | 29 |
| 6 | 1 3 DIMETHYLOROTIC ACID | 73 | 96% | 9% | 23 |
| 7 | OROTIDYLATE DECARBOXYLASE | 50 | 84% | 11% | 27 |
| 8 | CATALYTIC PROFICIENCY | 45 | 94% | 6% | 16 |
| 9 | OROTIDINE 5 MONOPHOSPHATE DECARBOXYLASE | 39 | 42% | 27% | 70 |
| 10 | OROTIDINE 5 PHOSPHATE DECARBOXYLASE | 20 | 35% | 18% | 47 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Catalytic proficiency: The unusual case of OMP decarboxylase | 2002 | 153 | 63 | 56% |
| OMP decarboxylase - An enigma persists | 2007 | 16 | 24 | 79% |
| Inhibition of orotidine 5 '-monophosphate decarboxylase and its therapeutic potential | 2008 | 9 | 43 | 72% |
| Insight into the catalytic mechanism of orotidine 5 '-phosphate decarboxylase from crystallography and mutagenesis | 2004 | 9 | 35 | 86% |
| On the importance of being zwitterionic: enzymatic catalysis of decarboxylation and deprotonation of cationic carbon | 2004 | 65 | 44 | 23% |
| Orotidine Monophosphate Decarboxylase - A Fascinating Workhorse Enzyme with Therapeutic Potential | 2015 | 0 | 76 | 86% |
| Enzymatic reactions involving novel mechanisms of carbanion stabilization | 2004 | 32 | 45 | 38% |
| What have theory and crystallography revealed about the mechanism of catalysis by orotidine monophosphate decarboxylase? | 2004 | 6 | 37 | 81% |
| Crystallographic studies of native and mutant orotidine 5 '-phosphate decarboxylases | 2004 | 1 | 25 | 96% |
| Catalysis by enzyme conformational change | 2004 | 21 | 50 | 40% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL DESIGN PREFORMULAT | 5 | 42% | 3.9% | 10 |
| 2 | MCLAUGHLIN ROTMAN UHN | 1 | 100% | 0.8% | 2 |
| 3 | PROTEIN ENGN NETWORK | 1 | 100% | 0.8% | 2 |
| 4 | THEORET PHYS CHEM GRP | 1 | 50% | 0.8% | 2 |
| 5 | GENIE CHIM BIOCHIM POLYTECH CLERMONT FERRAN | 1 | 50% | 0.4% | 1 |
| 6 | SRI SATYA SAI HIGHER LEARNING | 1 | 50% | 0.4% | 1 |
| 7 | TROP DIS UNIT INFECT DIS | 1 | 50% | 0.4% | 1 |
| 8 | MOL DESIGN PERFORMULAT | 0 | 33% | 0.4% | 1 |
| 9 | QUANTUM MOL ENGN | 0 | 33% | 0.4% | 1 |
| 10 | BIOMED ENVIRONM | 0 | 25% | 0.4% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000241915 | TRIOSEPHOSPHATE ISOMERASE//TRIOSEPHOSPHATE ISOMERASE DEFICIENCY//TRIOSEPHOSPHATE ISOMERASE TIM |
| 2 | 0.0000217003 | PHOSPHORIBOSYLTRANSFERASE//MOL PARASITOL DRUG DESIGN//HGXPRT |
| 3 | 0.0000201791 | MANDELATE RACEMASE//ENOLASE SUPERFAMILY//MANDELATE RACEMASE EC 5122 |
| 4 | 0.0000119880 | DIHYDROOROTASE//CARBAMOYL PHOSPHATE SYNTHETASE//ASPARTATE CARBAMOYLTRANSFERASE |
| 5 | 0.0000094712 | CIZ1//ENHANCER OF RUDIMENTARY//APICAL EXTRACELLULAR MATRIX |
| 6 | 0.0000085531 | EXTREMELY LOCALIZED MOLECULAR ORBITALS//ASEP MD//LINK ATOM |
| 7 | 0.0000084097 | TRANSKETOLASE//CARBOLIGATION//THIAMINE DIPHOSPHATE |
| 8 | 0.0000083262 | KETOSTEROID ISOMERASE//LOW BARRIER HYDROGEN BONDS//LOW BARRIER HYDROGEN BOND |
| 9 | 0.0000073857 | URIDINE PHOSPHORYLASE//5 PHENYLTHIOACYCLOURIDINE//ALPHA D RIBOSE 1 PHOSPHATE |
| 10 | 0.0000072573 | TRYPTOPHAN SYNTHASE//INDOLE 3 GLYCEROL PHOSPHATE SYNTHASE//SECT ENZYME STRUCT FUNCT |