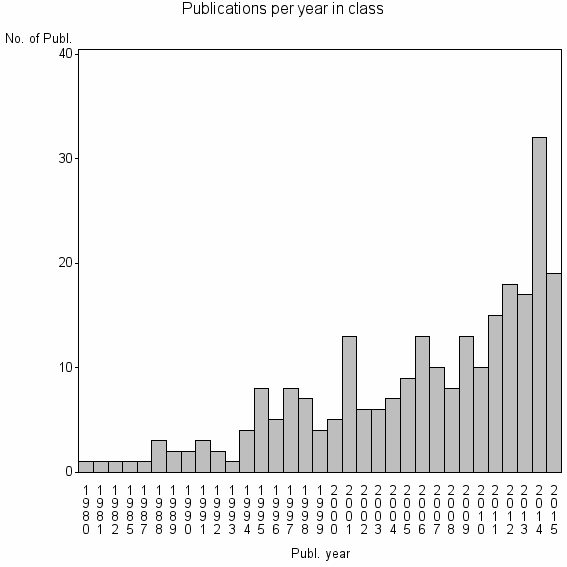
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 24669 | 255 | 35.9 | 78% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2410 | 3719 | HYDANTOINASE//D HYDANTOINASE//DIRECTED EVOLUTION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MANDELATE RACEMASE | Author keyword | 10 | 63% | 4% | 10 |
| 2 | ENOLASE SUPERFAMILY | Author keyword | 7 | 67% | 2% | 6 |
| 3 | MANDELATE RACEMASE EC 5122 | Author keyword | 6 | 100% | 2% | 4 |
| 4 | PSEUDOMONAS PUTIDA ATCC 12633 | Author keyword | 4 | 75% | 1% | 3 |
| 5 | N ACYLAMINO ACID RACEMASE | Author keyword | 4 | 56% | 2% | 5 |
| 6 | ENZYMATIC RACEMIZATION | Author keyword | 3 | 100% | 1% | 3 |
| 7 | D GALACTURONATE | Author keyword | 2 | 67% | 1% | 2 |
| 8 | NEW YORK SGX STRUCT GENOM | Address | 2 | 50% | 1% | 3 |
| 9 | N SUCCINYLAMINO ACID RACEMASE | Author keyword | 1 | 50% | 1% | 2 |
| 10 | O SUCCINYLBENZOATE SYNTHASE | Author keyword | 1 | 100% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ACID RACEMASE | 24 | 91% | 4% | 10 |
| 2 | ENOLASE SUPERFAMILY | 19 | 37% | 16% | 42 |
| 3 | O SUCCINYLBENZOATE SYNTHASE | 18 | 61% | 7% | 19 |
| 4 | MECHANISTIC PROPERTIES | 15 | 88% | 3% | 7 |
| 5 | ACYLAMINO ACID RACEMASE | 14 | 57% | 6% | 16 |
| 6 | D GLUCARATE DEHYDRATASE | 13 | 80% | 3% | 8 |
| 7 | AMYCOLATOPSIS SP TS 1 60 | 13 | 69% | 4% | 11 |
| 8 | MANDELATE RACEMASE | 12 | 31% | 13% | 34 |
| 9 | MUCONATE LACTONIZING ENZYME | 8 | 40% | 6% | 16 |
| 10 | AMIDOHYDROLASE SUPERFAMILY | 8 | 31% | 8% | 21 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Divergent Evolution in Enolase Superfamily: Strategies for Assigning Functions | 2012 | 36 | 34 | 74% |
| Divergent evolution in the enolase superfamily: the interplay of mechanism and specificity | 2005 | 110 | 48 | 77% |
| An evolutionary biochemist's perspective on promiscuity | 2015 | 5 | 42 | 12% |
| Thematic Minireview Series on Enzyme Evolution in the Post-genomic Era | 2012 | 5 | 1 | 100% |
| Evolution of enzyme superfamilies | 2006 | 97 | 36 | 25% |
| New Insights about Enzyme Evolution from Large Scale Studies of Sequence and Structure Relationships | 2014 | 3 | 59 | 29% |
| MANDELATE RACEMASE - STRUCTURE-FUNCTION STUDIES OF A PSEUDOSYMMETRIC ENZYME | 1995 | 71 | 25 | 60% |
| Divergent evolution of enzymatic function: Mechanistically diverse superfamilies and functionally distinct suprafamilies | 2001 | 275 | 138 | 13% |
| The substrate spectrum of mandelate racemase: Minimum structural requirements for substrates and substrate model | 2005 | 19 | 37 | 46% |
| Introduction to the Thematic Minireview Series on Enzyme Evolution | 2014 | 0 | 6 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NEW YORK SGX STRUCT GENOM | 2 | 50% | 1.2% | 3 |
| 2 | ABORATORY STRUCT BIOINFORMAT PROT DATA BA | 1 | 40% | 0.8% | 2 |
| 3 | LILLY BIOTECHNOL | 1 | 16% | 1.6% | 4 |
| 4 | BIOCHEM BIOPHYS CB 7260 | 1 | 50% | 0.4% | 1 |
| 5 | DEV PLICAT TECHNOL | 1 | 50% | 0.4% | 1 |
| 6 | GONOM | 1 | 50% | 0.4% | 1 |
| 7 | NAT PROD LINCHPIN | 1 | 25% | 0.8% | 2 |
| 8 | CHIROTECH TECHNOL | 0 | 20% | 0.8% | 2 |
| 9 | ALGORITHMS BIOINFORMAT | 0 | 25% | 0.4% | 1 |
| 10 | THERM VERFAHRENS UMWELTTECH | 0 | 25% | 0.4% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000275376 | BIOCATALYTIC PROMISCUITY//4 OXALOCROTONATE TAUTOMERASE//TAUTOMERASE SUPERFAMILY |
| 2 | 0.0000205996 | COMPLEX SCI SOFTWARE//3D ZERNIKE DESCRIPTOR//PROTEIN SURFACE SHAPE |
| 3 | 0.0000201791 | ODCASE//OROTIDINE 5 MONOPHOSPHATE DECARBOXYLASE//MOL DESIGN PREFORMULAT |
| 4 | 0.0000186040 | HYDANTOINASE//D HYDANTOINASE//L N CARBAMOYLASE |
| 5 | 0.0000167945 | RIBOKINASE//ADENOSINE KINASE//GLUCONATE DEHYDRATASE |
| 6 | 0.0000137864 | AMINOACYLASE//D AMINOACYLASE//L AMINOACYLASE |
| 7 | 0.0000113295 | ALANINE RACEMASE//MURD//GLUTAMATE RACEMASE |
| 8 | 0.0000103304 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY//JCSG//PROGRAM BIOINFORMAT SYST BIOL |
| 9 | 0.0000102484 | INOSITOL DEHYDROGENASE//INOSITOL CATABOLISM//MYO INOSITOL DEHYDROGENASE |
| 10 | 0.0000101284 | KETOSTEROID ISOMERASE//LOW BARRIER HYDROGEN BONDS//LOW BARRIER HYDROGEN BOND |