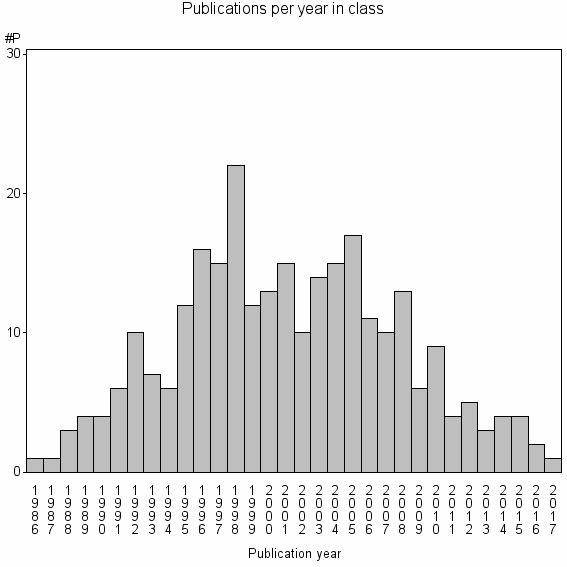
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 25676 | 275 | 37.6 | 88% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 33 | 3 | JOURNAL OF BACTERIOLOGY//MICROBIOLOGY//MOLECULAR MICROBIOLOGY | 107033 |
| 1115 | 2 | GENOME EDITING//CRISPR CAS9//CRISPR | 9706 |
| 25676 | 1 | GENE TRAP//GENE TRAPPING//SPLICING AROUND | 275 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GENE TRAP | authKW | 883722 | 13% | 22% | 37 |
| 2 | GENE TRAPPING | authKW | 444129 | 5% | 29% | 14 |
| 3 | SPLICING AROUND | authKW | 333112 | 1% | 100% | 3 |
| 4 | SPLICE ACCEPTOR TRAP | authKW | 222075 | 1% | 100% | 2 |
| 5 | ABHD2 | authKW | 148048 | 1% | 67% | 2 |
| 6 | SIGNAL SEQUENCE TRAP | authKW | 133235 | 2% | 20% | 6 |
| 7 | ALPHA BETA HYDROLASE DOMAIN CONTAINING 2 | authKW | 111037 | 0% | 100% | 1 |
| 8 | ALPHA BETA HYDROLASE FAMILY | authKW | 111037 | 0% | 100% | 1 |
| 9 | ALPHA BETA HYDROLASE PROTEIN | authKW | 111037 | 0% | 100% | 1 |
| 10 | BECKMAN B271 | address | 111037 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Developmental Biology | 2478 | 19% | 0% | 53 |
| 2 | Biochemistry & Molecular Biology | 767 | 45% | 0% | 123 |
| 3 | Genetics & Heredity | 677 | 21% | 0% | 59 |
| 4 | Cell Biology | 587 | 25% | 0% | 68 |
| 5 | Biochemical Research Methods | 357 | 13% | 0% | 35 |
| 6 | Anatomy & Morphology | 355 | 5% | 0% | 14 |
| 7 | Biotechnology & Applied Microbiology | 324 | 16% | 0% | 43 |
| 8 | Biophysics | 47 | 6% | 0% | 16 |
| 9 | Biology | 23 | 3% | 0% | 9 |
| 10 | Virology | 13 | 2% | 0% | 6 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BECKMAN B271 | 111037 | 0% | 100% | 1 |
| 2 | DOCTEUR P TAM | 111037 | 0% | 100% | 1 |
| 3 | INGOLSTAEDTER | 111037 | 0% | 100% | 1 |
| 4 | INVEST SCITISSUE ENGN | 111037 | 0% | 100% | 1 |
| 5 | MOL GENET BIOFUNCT BIOMED SCI | 111037 | 0% | 100% | 1 |
| 6 | TECH EVALUAT EUROPE | 111037 | 0% | 100% | 1 |
| 7 | UR 101IFRBAIM | 111037 | 0% | 100% | 1 |
| 8 | MAX DELBRUCK CARL VON LINNE WEG 10 | 111035 | 1% | 50% | 2 |
| 9 | BIO OURCE PROJECT NBRP | 55518 | 0% | 50% | 1 |
| 10 | MACDONALD MED S | 55518 | 0% | 50% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | TRANSGENIC RESEARCH | 4252 | 3% | 0% | 8 |
| 2 | DEVELOPMENTAL DYNAMICS | 4024 | 5% | 0% | 13 |
| 3 | BIOTECHFORUM ( BTF ) | 2093 | 3% | 0% | 7 |
| 4 | GENESIS | 1831 | 2% | 0% | 5 |
| 5 | NUCLEIC ACIDS RESEARCH | 1217 | 8% | 0% | 21 |
| 6 | ALCOHOL RESEARCH-CURRENT REVIEWS | 1131 | 0% | 1% | 1 |
| 7 | DEVELOPMENT GROWTH & DIFFERENTIATION | 1084 | 2% | 0% | 5 |
| 8 | CURRENT TOPICS IN DEVELOPMENTAL BIOLOGY | 974 | 1% | 0% | 3 |
| 9 | METHODS IN ENZYMOLOGY | 966 | 4% | 0% | 12 |
| 10 | NATURE BIOTECHNOLOGY | 671 | 1% | 0% | 4 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | FRIEDEL, RH , SORIANO, P , (2010) GENE TRAP MUTAGENESIS IN THE MOUSE.METHODS IN ENZYMOLOGY, VOL 477: GUIDE TO TECHNIQUES IN MOUSE DEVELOPMENT, PART B: MOUSE MOLECULAR GENETICS, SECOND EDITION. VOL. 477. ISSUE . P. 243-269 | 47 | 57% | 16 |
| 2 | BRICKMAN, JM , TSAKIRIDIS, A , TO, C , STANFORD, WL , (2010) A WIDER CONTEXT FOR GENE TRAP MUTAGENESIS.METHODS IN ENZYMOLOGY, VOL 477: GUIDE TO TECHNIQUES IN MOUSE DEVELOPMENT, PART B: MOUSE MOLECULAR GENETICS, SECOND EDITION. VOL. 477. ISSUE . P. 271 -295 | 41 | 54% | 7 |
| 3 | SKARNES, WC , (2000) GENE TRAPPING METHODS FOR THE IDENTIFICATION AND FUNCTIONAL ANALYSIS OF CELL SURFACE PROTEINS IN MICE.APPLICATIONS OF CHIMERIC GENES AND HYBRID PROTEINS, PT C. VOL. 328. ISSUE . P. 592 -615 | 27 | 90% | 57 |
| 4 | STANFORD, WL , COHN, JB , CORDES, SP , (2001) GENE-TRAP MUTAGENESIS: PAST, PRESENT AND BEYOND.NATURE REVIEWS GENETICS. VOL. 2. ISSUE 10. P. 756-768 | 37 | 40% | 207 |
| 5 | STANFORD, WL , EPP, T , REID, T , ROSSANT, J , (2006) GENE TRAPPING IN EMBRYONIC STEM CELLS.STEM CELL TOOLS AND OTHER EXPERIMENTAL PROTOCOLS. VOL. 420. ISSUE . P. 136-162 | 23 | 61% | 14 |
| 6 | KITAJIMA, K , TAKEUCHI, T , (1998) MOUSE GENE TRAP APPROACH: IDENTIFICATION OF NOVEL GENES AND CHARACTERIZATION OF THEIR BIOLOGICAL FUNCTIONS.BIOCHEMISTRY AND CELL BIOLOGY-BIOCHIMIE ET BIOLOGIE CELLULAIRE. VOL. 76. ISSUE 6. P. 1029 -1037 | 37 | 48% | 0 |
| 7 | FORRAI, A , ROBB, L , (2005) THE GENE TRAP RESOURCE: A TREASURE TROVE FOR HEMOPOIESIS RESEARCH.EXPERIMENTAL HEMATOLOGY. VOL. 33. ISSUE 8. P. 845-856 | 46 | 30% | 8 |
| 8 | CAMUS, A , BARRA, J , BABINET, C , (1998) GENE TRAP STRATEGIES FOR THE IDENTIFICATION OF DEVELOPMENTALLY IMPORTANT MURINE GENES.M S-MEDECINE SCIENCES. VOL. 14. ISSUE 11. P. 1157 -1166 | 30 | 53% | 0 |
| 9 | YAMAMURA, K , ARAKI, K , (2008) GENE TRAP MUTAGENESIS IN MICE: NEW PERSPECTIVES AND TOOLS IN CANCER RESEARCH.CANCER SCIENCE. VOL. 99. ISSUE 1. P. 1 -6 | 25 | 42% | 4 |
| 10 | KUHNERT, F , STUHLMANN, H , (2004) IDENTIFYING EARLY VASCULAR GENES THROUGH GENE TRAPPING IN MOUSE EMBRYONIC STEM CELLS.DEVELOPMENTAL VASCULAR BIOLOGY. VOL. 62. ISSUE . P. 261 -+ | 26 | 38% | 4 |
Classes with closest relation at Level 1 |