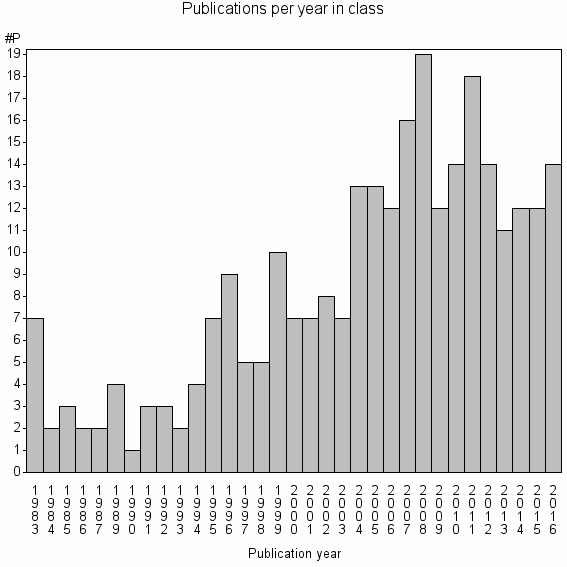
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 25557 | 278 | 41.1 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 271 | 3 | MITOCHONDRIA//MITOCHONDRIAL GENOME//MITOCHONDRIAL DNA PART A | 43930 |
| 976 | 2 | MITOCHONDRIAL GENOME//MITOCHONDRIAL DNA PART A//MITOCHONDRIAL DNA | 10638 |
| 25557 | 1 | NUMT//NUMTS//MITOCHONDRIAL PSEUDOGENES | 278 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NUMT | authKW | 2820694 | 15% | 60% | 43 |
| 2 | NUMTS | authKW | 1594674 | 10% | 52% | 28 |
| 3 | MITOCHONDRIAL PSEUDOGENES | authKW | 494273 | 2% | 75% | 6 |
| 4 | NUCLEAR MITOCHONDRIAL PSEUDOGENES | authKW | 494273 | 2% | 75% | 6 |
| 5 | NUMTDNA | authKW | 439356 | 1% | 100% | 4 |
| 6 | NUCLEAR INSERTION | authKW | 351484 | 1% | 80% | 4 |
| 7 | NUPTS | authKW | 251057 | 1% | 57% | 4 |
| 8 | NUCLEAR COPIES | authKW | 219678 | 1% | 100% | 2 |
| 9 | NUCLEAR MTDNA INSERTIONS | authKW | 219678 | 1% | 100% | 2 |
| 10 | NUCLEAR INTEGRATION | authKW | 197708 | 1% | 60% | 3 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 5159 | 56% | 0% | 156 |
| 2 | Evolutionary Biology | 4477 | 27% | 0% | 74 |
| 3 | Biochemistry & Molecular Biology | 701 | 43% | 0% | 119 |
| 4 | Cell Biology | 107 | 12% | 0% | 33 |
| 5 | Biotechnology & Applied Microbiology | 90 | 9% | 0% | 25 |
| 6 | Ecology | 74 | 7% | 0% | 19 |
| 7 | Entomology | 37 | 3% | 0% | 9 |
| 8 | Biology | 29 | 4% | 0% | 10 |
| 9 | Zoology | 24 | 4% | 0% | 11 |
| 10 | Plant Sciences | 13 | 5% | 0% | 13 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CNRSUNITE GENET BIOCHIM DEV | 109839 | 0% | 100% | 1 |
| 2 | CNRSURA1300UNITE GENET MOL LEVU | 109839 | 0% | 100% | 1 |
| 3 | ESTABLISSEMENT FRANCAIS SANG BRETAGNE | 109839 | 0% | 100% | 1 |
| 4 | GENET EPIGENET ALTERAT GENOME GRP GENEPI | 109839 | 0% | 100% | 1 |
| 5 | INSERMU613 SANTE RECH MED | 109839 | 0% | 100% | 1 |
| 6 | INTRAMURAL SUPPORT PROGRAM GENOM ERS | 109839 | 0% | 100% | 1 |
| 7 | LEHRSTUHL MOL BIOL PFLANZEN | 109839 | 0% | 100% | 1 |
| 8 | MITOCHONDRIAL GERONTOL AGE RELATED DIS GRP | 109839 | 0% | 100% | 1 |
| 9 | MOL GENET GEMO | 109839 | 0% | 100% | 1 |
| 10 | MOL PLANT PHYSIOL BIOTECHNOL CGB | 109839 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR BIOLOGY AND EVOLUTION | 8678 | 8% | 0% | 21 |
| 2 | MITOCHONDRION | 6603 | 3% | 1% | 8 |
| 3 | JOURNAL OF ZOOLOGICAL SYSTEMATICS AND EVOLUTIONARY RESEARCH | 2565 | 1% | 1% | 4 |
| 4 | FUNCTIONAL & INTEGRATIVE GENOMICS | 2024 | 1% | 1% | 3 |
| 5 | JOURNAL OF MOLECULAR EVOLUTION | 1834 | 3% | 0% | 8 |
| 6 | GENOME BIOLOGY AND EVOLUTION | 1296 | 1% | 0% | 4 |
| 7 | PLOS GENETICS | 883 | 3% | 0% | 7 |
| 8 | RUSSIAN JOURNAL OF GENETICS | 795 | 2% | 0% | 5 |
| 9 | GENOME RESEARCH | 713 | 1% | 0% | 3 |
| 10 | INSECT MOLECULAR BIOLOGY | 685 | 1% | 0% | 3 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NUMT | 2820694 | 15% | 60% | 43 | Search NUMT | Search NUMT |
| 2 | NUMTS | 1594674 | 10% | 52% | 28 | Search NUMTS | Search NUMTS |
| 3 | MITOCHONDRIAL PSEUDOGENES | 494273 | 2% | 75% | 6 | Search MITOCHONDRIAL+PSEUDOGENES | Search MITOCHONDRIAL+PSEUDOGENES |
| 4 | NUCLEAR MITOCHONDRIAL PSEUDOGENES | 494273 | 2% | 75% | 6 | Search NUCLEAR+MITOCHONDRIAL+PSEUDOGENES | Search NUCLEAR+MITOCHONDRIAL+PSEUDOGENES |
| 5 | NUMTDNA | 439356 | 1% | 100% | 4 | Search NUMTDNA | Search NUMTDNA |
| 6 | NUCLEAR INSERTION | 351484 | 1% | 80% | 4 | Search NUCLEAR+INSERTION | Search NUCLEAR+INSERTION |
| 7 | NUPTS | 251057 | 1% | 57% | 4 | Search NUPTS | Search NUPTS |
| 8 | NUCLEAR COPIES | 219678 | 1% | 100% | 2 | Search NUCLEAR+COPIES | Search NUCLEAR+COPIES |
| 9 | NUCLEAR MTDNA INSERTIONS | 219678 | 1% | 100% | 2 | Search NUCLEAR+MTDNA+INSERTIONS | Search NUCLEAR+MTDNA+INSERTIONS |
| 10 | NUCLEAR INTEGRATION | 197708 | 1% | 60% | 3 | Search NUCLEAR+INTEGRATION | Search NUCLEAR+INTEGRATION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HAZKANI-COVO, E , ZELLER, RM , MARTIN, W , (2010) MOLECULAR POLTERGEISTS: MITOCHONDRIAL DNA COPIES (NUMTS) IN SEQUENCED NUCLEAR GENOMES.PLOS GENETICS. VOL. 6. ISSUE 2. P. - | 49 | 59% | 152 |
| 2 | SONG, HJ , MOULTON, MJ , WHITING, MF , (2014) RAMPANT NUCLEAR INSERTION OF MTDNA ACROSS DIVERSE LINEAGES WITHIN ORTHOPTERA (INSECTA).PLOS ONE. VOL. 9. ISSUE 10. P. - | 47 | 59% | 6 |
| 3 | BENSASSON, D , ZHANG, DX , HARTL, DL , HEWITT, GM , (2001) MITOCHONDRIAL PSEUDOGENES: EVOLUTION'S MISPLACED WITNESSES.TRENDS IN ECOLOGY & EVOLUTION. VOL. 16. ISSUE 6. P. 314 -321 | 35 | 65% | 598 |
| 4 | SHI, HZ , DONG, J , IRWIN, DM , ZHANG, SY , MAO, XG , (2016) REPETITIVE TRANSPOSITIONS OF MITOCHONDRIAL DNA SEQUENCES TO THE NUCLEUS DURING THE RADIATION OF HORSESHOE BATS (RHINOLOPHUS, CHIROPTERA).GENE. VOL. 581. ISSUE 2. P. 161 -169 | 26 | 67% | 0 |
| 5 | BEHURA, SK , (2012) RARITY OF TAGSNPS ACROSS TRANSFERRED MTDNA INSERTS IN HUMAN GENOME.MOLECULAR BIOLOGY. VOL. 46. ISSUE 1. P. 102-106 | 19 | 86% | 0 |
| 6 | SACERDOT, C , CASAREGOLA, S , LAFONTAINE, I , TEKAIA, F , DUJON, B , OZIER-KALOGEROPOULOS, O , (2008) PROMISCUOUS DNA IN THE NUCLEAR GENOMES OF HEMIASCOMYCETOUS YEASTS.FEMS YEAST RESEARCH. VOL. 8. ISSUE 6. P. 846 -857 | 24 | 73% | 15 |
| 7 | LEISTER, D , (2005) ORIGIN, EVOLUTION AND GENETIC EFFECTS OF NUCLEAR INSERTIONS OF ORGANELLE DNA.TRENDS IN GENETICS. VOL. 21. ISSUE 12. P. 655 -663 | 34 | 50% | 84 |
| 8 | ANTUNES, A , PONTIUS, J , RAMOS, MJ , O'BRIEN, SJ , JOHNSON, WE , (2007) MITOCHONDRIAL INTROGRESSIONS INTO THE NUCLEAR GENOME OF THE DOMESTIC CAT.JOURNAL OF HEREDITY. VOL. 98. ISSUE 5. P. 414-420 | 24 | 73% | 14 |
| 9 | SONG, HJ , MOULTON, MJ , HIATT, KD , WHITING, MF , (2013) UNCOVERING HISTORICAL SIGNATURE OF MITOCHONDRIAL DNA HIDDEN IN THE NUCLEAR GENOME: THE BIOGEOGRAPHY OF SCHISTOCERCA REVISITED.CLADISTICS. VOL. 29. ISSUE 6. P. 643-662 | 34 | 43% | 3 |
| 10 | JENSEN-SEAMAN, MI , WILDSCHUTTE, JH , SOTO-CALDERON, ID , ANTHONY, NM , (2009) A COMPARATIVE APPROACH SHOWS DIFFERENCES IN PATTERNS OF NUMT INSERTION DURING HOMINOID EVOLUTION.JOURNAL OF MOLECULAR EVOLUTION. VOL. 68. ISSUE 6. P. 688 -699 | 31 | 50% | 17 |
Classes with closest relation at Level 1 |