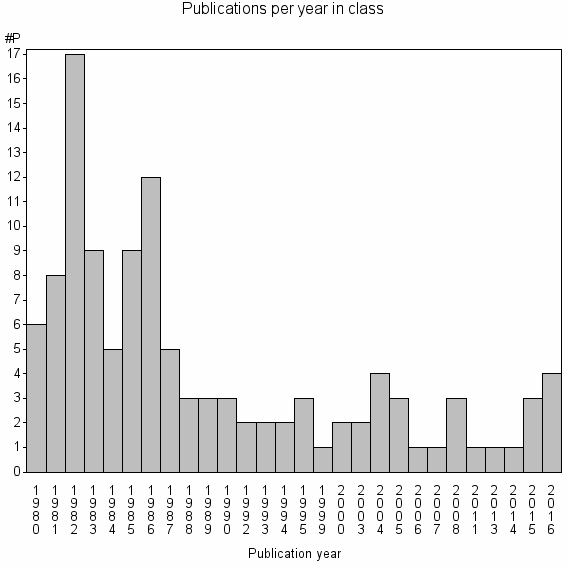
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 34076 | 116 | 35.2 | 26% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 260 | 2 | PROTEIN FOLDING//JOURNAL OF CHEMICAL THEORY AND COMPUTATION//JOURNAL OF COMPUTATIONAL CHEMISTRY | 19582 |
| 34076 | 1 | UNIVERSAL EXCHANGE LANGUAGE//MEDICAL INFERENCE//BIOINFORMATICS MESSAGING | 116 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | UNIVERSAL EXCHANGE LANGUAGE | authKW | 789714 | 3% | 100% | 3 |
| 2 | MEDICAL INFERENCE | authKW | 592284 | 3% | 75% | 3 |
| 3 | BIOINFORMATICS MESSAGING | authKW | 263238 | 1% | 100% | 1 |
| 4 | BIOSTAT EPIDEMIOL EVIDENCE BASED MED | address | 263238 | 1% | 100% | 1 |
| 5 | CLINICAL MESSAGING | authKW | 263238 | 1% | 100% | 1 |
| 6 | COMPLEX DATA SETS | authKW | 263238 | 1% | 100% | 1 |
| 7 | DNA MARK UP | authKW | 263238 | 1% | 100% | 1 |
| 8 | GENOMIC MESSAGING SYSTEM | authKW | 263238 | 1% | 100% | 1 |
| 9 | GENOMICS LANGUAGE | authKW | 263238 | 1% | 100% | 1 |
| 10 | HEALTHMINER R | authKW | 263238 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Nutrition & Dietetics | 426 | 16% | 0% | 19 |
| 2 | Mathematical & Computational Biology | 295 | 9% | 0% | 11 |
| 3 | Computer Science, Interdisciplinary Applications | 183 | 11% | 0% | 13 |
| 4 | Biology | 165 | 11% | 0% | 13 |
| 5 | Engineering, Biomedical | 75 | 7% | 0% | 8 |
| 6 | Biochemistry & Molecular Biology | 60 | 22% | 0% | 26 |
| 7 | Biophysics | 52 | 9% | 0% | 10 |
| 8 | Biochemical Research Methods | 38 | 7% | 0% | 8 |
| 9 | Physics, Atomic, Molecular & Chemical | 17 | 6% | 0% | 7 |
| 10 | Chemistry, Multidisciplinary | 15 | 10% | 0% | 12 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOSTAT EPIDEMIOL EVIDENCE BASED MED | 263238 | 1% | 100% | 1 |
| 2 | IBM LIFE SCI | 263238 | 1% | 100% | 1 |
| 3 | MD GR E | 263238 | 1% | 100% | 1 |
| 4 | ST MATTHEWS UNIV MED | 263238 | 1% | 100% | 1 |
| 5 | ENVIRONM SPECIMEN BANK HUMAN TISSUES | 43871 | 1% | 17% | 1 |
| 6 | GLOBAL COMPOUND SCI | 32903 | 1% | 13% | 1 |
| 7 | MODERN BIOL | 17258 | 2% | 3% | 2 |
| 8 | CHEM QUANTUM THEORY PROJECT | 10123 | 1% | 4% | 1 |
| 9 | THOMAS J WATSON | 1957 | 5% | 0% | 6 |
| 10 | SCI CHEM BUNKYO KU | 784 | 1% | 0% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | WATER SCIENCE REVIEWS | 43871 | 1% | 17% | 1 |
| 2 | COMPUTERS IN BIOLOGY AND MEDICINE | 5775 | 7% | 0% | 8 |
| 3 | ARCHIVOS LATINOAMERICANOS DE NUTRICION | 4005 | 4% | 0% | 5 |
| 4 | JOURNAL OF PROTEOME RESEARCH | 2757 | 7% | 0% | 8 |
| 5 | CRC CRITICAL REVIEWS IN FOOD SCIENCE AND NUTRITION | 2055 | 1% | 1% | 1 |
| 6 | CRC CRITICAL REVIEWS IN BIOCHEMISTRY | 1852 | 1% | 1% | 1 |
| 7 | BIOCHEMICAL SOCIETY SYMPOSIA | 1421 | 1% | 1% | 1 |
| 8 | JOURNAL OF MOLECULAR GRAPHICS | 1195 | 1% | 0% | 1 |
| 9 | ADVANCES IN MICROBIAL PHYSIOLOGY | 1128 | 1% | 0% | 1 |
| 10 | SPECULATIONS IN SCIENCE AND TECHNOLOGY | 948 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | UNIVERSAL EXCHANGE LANGUAGE | 789714 | 3% | 100% | 3 | Search UNIVERSAL+EXCHANGE+LANGUAGE | Search UNIVERSAL+EXCHANGE+LANGUAGE |
| 2 | MEDICAL INFERENCE | 592284 | 3% | 75% | 3 | Search MEDICAL+INFERENCE | Search MEDICAL+INFERENCE |
| 3 | BIOINFORMATICS MESSAGING | 263238 | 1% | 100% | 1 | Search BIOINFORMATICS+MESSAGING | Search BIOINFORMATICS+MESSAGING |
| 4 | CLINICAL MESSAGING | 263238 | 1% | 100% | 1 | Search CLINICAL+MESSAGING | Search CLINICAL+MESSAGING |
| 5 | COMPLEX DATA SETS | 263238 | 1% | 100% | 1 | Search COMPLEX+DATA+SETS | Search COMPLEX+DATA+SETS |
| 6 | DNA MARK UP | 263238 | 1% | 100% | 1 | Search DNA+MARK+UP | Search DNA+MARK+UP |
| 7 | GENOMIC MESSAGING SYSTEM | 263238 | 1% | 100% | 1 | Search GENOMIC+MESSAGING+SYSTEM | Search GENOMIC+MESSAGING+SYSTEM |
| 8 | GENOMICS LANGUAGE | 263238 | 1% | 100% | 1 | Search GENOMICS+LANGUAGE | Search GENOMICS+LANGUAGE |
| 9 | HEALTHMINER R | 263238 | 1% | 100% | 1 | Search HEALTHMINER+R | Search HEALTHMINER+R |
| 10 | HYPERBOLIC DIRAC NET | 263238 | 1% | 100% | 1 | Search HYPERBOLIC+DIRAC+NET | Search HYPERBOLIC+DIRAC+NET |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ROBSON, B , (2008) CLINICAL AND PHARMACOGENOMIC DATA MINING: 4. THE FANO PROGRAM AND COMMAND SET AS AN EXAMPLE OF TOOLS FOR BIOMEDICAL DISCOVERY AND EVIDENCE BASED MEDICINE.JOURNAL OF PROTEOME RESEARCH. VOL. 7. ISSUE 9. P. 3922 -3947 | 9 | 75% | 1 |
| 2 | ROBSON, B , CARUSO, TP , BALIS, UGJ , (2013) SUGGESTIONS FOR A WEB BASED UNIVERSAL EXCHANGE AND INFERENCE LANGUAGE FOR MEDICINE.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 43. ISSUE 12. P. 2297 -2310 | 6 | 86% | 1 |
| 3 | ROBSON, B , (2016) STUDIES IN USING A UNIVERSAL EXCHANGE AND INFERENCE LANGUAGE FOR EVIDENCE BASED MEDICINE. SEMI-AUTOMATED LEARNING AND REASONING FOR PICO METHODOLOGY, SYSTEMATIC REVIEW, AND ENVIRONMENTAL EPIDEMIOLOGY.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 79. ISSUE . P. 299 -323 | 9 | 53% | 0 |
| 4 | ROBSON, B , (2014) HYPERBOLIC DIRAC NETS FOR MEDICAL DECISION SUPPORT. THEORY, METHODS, AND COMPARISON WITH BAYES NETS.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 51. ISSUE . P. 183 -197 | 6 | 75% | 1 |
| 5 | ROBSON, B , BORAY, S , (2016) STUDIES OF THE ROLE OF A SMART WEB FOR PRECISION MEDICINE SUPPORTED BY BIOBANKING.PERSONALIZED MEDICINE. VOL. 13. ISSUE 4. P. 361 -380 | 8 | 53% | 0 |
| 6 | ROBSON, B , BORAY, S , (2015) IMPLEMENTATION OF A WEB BASED UNIVERSAL EXCHANGE AND INFERENCE LANGUAGE FOR MEDICINE: SPARSE DATA, PROBABILITIES AND INFERENCE IN DATA MINING OF CLINICAL DATA REPOSITORIES.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 66. ISSUE . P. 82 -102 | 9 | 47% | 0 |
| 7 | ROBSON, B , MUSHLIN, R , (2005) GENOMIC MESSAGING SYSTEM LANGUAGE INCLUDING COMMAND EXTENSIONS FOR CLINICAL DATA CATEGORIES.JOURNAL OF PROTEOME RESEARCH. VOL. 4. ISSUE 2. P. 275 -299 | 5 | 83% | 3 |
| 8 | ROBSON, B , CARUSO, TP , BALIS, UGJ , (2015) SUGGESTIONS FOR A WEB BASED UNIVERSAL EXCHANGE AND INFERENCE LANGUAGE FOR MEDICINE. CONTINUITY OF PATIENT CARE WITH PCAST DISAGGREGATION.COMPUTERS IN BIOLOGY AND MEDICINE. VOL. 56. ISSUE . P. 51 -66 | 6 | 33% | 1 |
| 9 | ROBSON, B , (2004) THE DRAGON ON THE GOLD: MYTHS AND REALITIES FOR DATA MINING IN BIOMEDICINE AND BIOTECHNOLOGY USING DIGITAL AND MOLECULAR LIBRARIES.JOURNAL OF PROTEOME RESEARCH. VOL. 3. ISSUE 6. P. 1113-1119 | 2 | 100% | 8 |
| 10 | FINNEY, JL , GOODFELLOW, JM , HOWELL, PL , VOVELLE, F , (1985) COMPUTER-SIMULATION OF AQUEOUS BIOMOLECULAR SYSTEMS.JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS. VOL. 3. ISSUE 3. P. 599-622 | 8 | 47% | 20 |
Classes with closest relation at Level 1 |