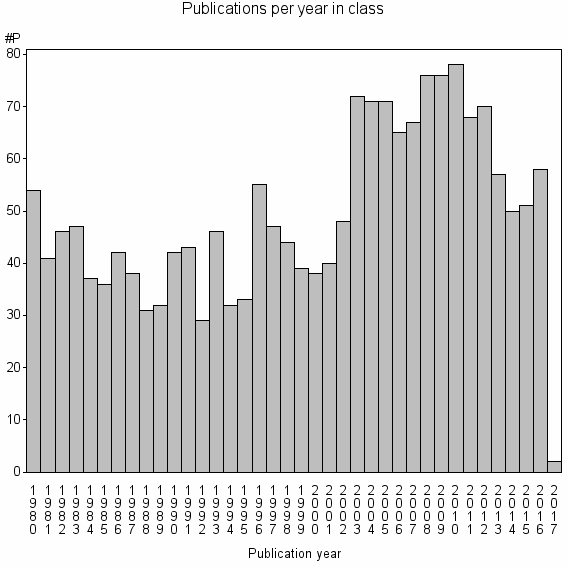
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3291 | 1872 | 33.8 | 64% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 236 | 3 | ADENOSINE//NUCLEOSIDES NUCLEOTIDES & NUCLEIC ACIDS//ADENOSINE RECEPTORS | 48331 |
| 3291 | 2 | PURINE NUCLEOSIDE PHOSPHORYLASE//CARBOHYDRATE CHEM TEAM//METHIONINE DEPENDENCE | 1872 |
| 9868 | 1 | PURINE NUCLEOSIDE PHOSPHORYLASE//CARBOHYDRATE CHEM TEAM//NUCLEOSIDE HYDROLASE | 1113 |
| 22214 | 1 | MTAP//METHYLTHIOADENOSINE PHOSPHORYLASE//METHIONINE SALVAGE PATHWAY | 390 |
| 22761 | 1 | METHIONINE DEPENDENCE//METHIONINASE//METHIONINE GAMMA LYASE | 369 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PURINE NUCLEOSIDE PHOSPHORYLASE | authKW | 1105184 | 7% | 53% | 127 |
| 2 | CARBOHYDRATE CHEM TEAM | address | 350619 | 2% | 50% | 43 |
| 3 | METHIONINE DEPENDENCE | authKW | 277268 | 1% | 100% | 17 |
| 4 | NUCLEOSIDE HYDROLASE | authKW | 262491 | 1% | 62% | 26 |
| 5 | NUCLEOSIDE PHOSPHORYLASE | authKW | 257496 | 2% | 53% | 30 |
| 6 | METHIONINASE | authKW | 251633 | 1% | 86% | 18 |
| 7 | BIOQUIM ESTRUTURAL | address | 220174 | 1% | 75% | 18 |
| 8 | METHYLTHIOADENOSINE PHOSPHORYLASE | authKW | 177590 | 1% | 78% | 14 |
| 9 | MTAP | authKW | 162522 | 1% | 59% | 17 |
| 10 | INVEST BIOTECNOL SUSTENTABLE LIBIOS | address | 132108 | 0% | 90% | 9 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 5103 | 44% | 0% | 829 |
| 2 | Biophysics | 1362 | 11% | 0% | 200 |
| 3 | Oncology | 830 | 13% | 0% | 239 |
| 4 | Chemistry, Medicinal | 629 | 6% | 0% | 115 |
| 5 | Biochemical Research Methods | 532 | 7% | 0% | 122 |
| 6 | Biotechnology & Applied Microbiology | 479 | 8% | 0% | 153 |
| 7 | Chemistry, Organic | 327 | 8% | 0% | 142 |
| 8 | Pharmacology & Pharmacy | 150 | 8% | 0% | 148 |
| 9 | Crystallography | 98 | 3% | 0% | 49 |
| 10 | Microbiology | 85 | 4% | 0% | 81 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CARBOHYDRATE CHEM TEAM | 350619 | 2% | 50% | 43 |
| 2 | BIOQUIM ESTRUTURAL | 220174 | 1% | 75% | 18 |
| 3 | INVEST BIOTECNOL SUSTENTABLE LIBIOS | 132108 | 0% | 90% | 9 |
| 4 | DIPARTIMENTO BIOCHIM BIOFIS F CEDRANGOLO | 102074 | 1% | 48% | 13 |
| 5 | PROGRAMA POS MED CIENCIAS SAUDE | 83945 | 1% | 23% | 22 |
| 6 | REDE BRASILEIRA PESQUISAS TB | 73391 | 0% | 75% | 6 |
| 7 | ITALIAN BIOCATALYSIS | 52832 | 0% | 36% | 9 |
| 8 | BIOCRYSTALLOG UNIT | 49696 | 0% | 38% | 8 |
| 9 | STRUCT BIOL ZOOCHEM CEIS | 48930 | 0% | 100% | 3 |
| 10 | PESQUISAS BIOL MOL FUNC | 39788 | 1% | 20% | 12 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEOSIDES NUCLEOTIDES & NUCLEIC ACIDS | 5535 | 2% | 1% | 40 |
| 2 | AGRICULTURAL AND BIOLOGICAL CHEMISTRY | 2973 | 2% | 1% | 36 |
| 3 | BIOCHEMISTRY | 2746 | 5% | 0% | 97 |
| 4 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY | 1756 | 1% | 1% | 22 |
| 5 | CURRENT DRUG TARGETS | 1726 | 1% | 1% | 14 |
| 6 | NUCLEOSIDES & NUCLEOTIDES | 1188 | 0% | 1% | 7 |
| 7 | JOURNAL OF CARBOHYDRATES-NUCLEOSIDES-NUCLEOTIDES | 1121 | 0% | 3% | 2 |
| 8 | BIOCATALYSIS AND BIOTRANSFORMATION | 859 | 0% | 1% | 7 |
| 9 | BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS | 854 | 1% | 0% | 13 |
| 10 | CANCER RESEARCH | 848 | 3% | 0% | 47 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PURINE NUCLEOSIDE PHOSPHORYLASE | 1105184 | 7% | 53% | 127 | Search PURINE+NUCLEOSIDE+PHOSPHORYLASE | Search PURINE+NUCLEOSIDE+PHOSPHORYLASE |
| 2 | METHIONINE DEPENDENCE | 277268 | 1% | 100% | 17 | Search METHIONINE+DEPENDENCE | Search METHIONINE+DEPENDENCE |
| 3 | NUCLEOSIDE HYDROLASE | 262491 | 1% | 62% | 26 | Search NUCLEOSIDE+HYDROLASE | Search NUCLEOSIDE+HYDROLASE |
| 4 | NUCLEOSIDE PHOSPHORYLASE | 257496 | 2% | 53% | 30 | Search NUCLEOSIDE+PHOSPHORYLASE | Search NUCLEOSIDE+PHOSPHORYLASE |
| 5 | METHIONINASE | 251633 | 1% | 86% | 18 | Search METHIONINASE | Search METHIONINASE |
| 6 | METHYLTHIOADENOSINE PHOSPHORYLASE | 177590 | 1% | 78% | 14 | Search METHYLTHIOADENOSINE+PHOSPHORYLASE | Search METHYLTHIOADENOSINE+PHOSPHORYLASE |
| 7 | MTAP | 162522 | 1% | 59% | 17 | Search MTAP | Search MTAP |
| 8 | METHIONINE GAMMA LYASE | 131246 | 1% | 62% | 13 | Search METHIONINE+GAMMA+LYASE | Search METHIONINE+GAMMA+LYASE |
| 9 | L METHIONINE GAMMA LYASE | 120097 | 0% | 82% | 9 | Search L+METHIONINE+GAMMA+LYASE | Search L+METHIONINE+GAMMA+LYASE |
| 10 | METHIONINE SALVAGE PATHWAY | 117421 | 1% | 60% | 12 | Search METHIONINE+SALVAGE+PATHWAY | Search METHIONINE+SALVAGE+PATHWAY |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | BZOWSKA, A , KULIKOWSKA, E , SHUGAR, D , (2000) PURINE NUCLEOSIDE PHOSPHORYLASES: PROPERTIES, FUNCTIONS, AND CLINICAL ASPECTS.PHARMACOLOGY & THERAPEUTICS. VOL. 88. ISSUE 3. P. 349 -425 | 145 | 49% | 270 |
| 2 | SILVA, RG , NUNES, JES , CANDURI, F , BORGES, JC , GAVA, LM , MORENO, FB , BASSO, LA , SANTOS, DS , (2007) PURINE NUCLEOSIDE PHOSPHORYLASE: A POTENTIAL TARGET FOR THE DEVELOPMENT OF DRUGS TO TREAT T-CELL- AND APICOMPLEXAN PARASITE-MEDIATED DISEASES.CURRENT DRUG TARGETS. VOL. 8. ISSUE 3. P. 413 -422 | 65 | 86% | 12 |
| 3 | EVANS, GB , SCHRAMM, VL , TYLER, PC , (2015) THE IMMUCILLINS: DESIGN, SYNTHESIS AND APPLICATION OF TRANSITION-STATE ANALOGUES.CURRENT MEDICINAL CHEMISTRY. VOL. 22. ISSUE 34. P. 3897 -3909 | 48 | 86% | 0 |
| 4 | HOFFMAN, RM , (2015) DEVELOPMENT OF RECOMBINANT METHIONINASE TO TARGET THE GENERAL CANCER-SPECIFIC METABOLIC DEFECT OF METHIONINE DEPENDENCE: A 40-YEAR ODYSSEY.EXPERT OPINION ON BIOLOGICAL THERAPY. VOL. 15. ISSUE 1. P. 21 -31 | 39 | 91% | 5 |
| 5 | CAVUOTO, P , FENECH, MF , (2012) A REVIEW OF METHIONINE DEPENDENCY AND THE ROLE OF METHIONINE RESTRICTION IN CANCER GROWTH CONTROL AND LIFE-SPAN EXTENSION.CANCER TREATMENT REVIEWS. VOL. 38. ISSUE 6. P. 726-736 | 58 | 56% | 50 |
| 6 | DURANDO, X , THIVAT, E , GIMBERGUES, P , CELLARIER, E , ABRIAL, C , DIB, M , TACCA, O , CHOLLET, P , (2008) METHIONINE DEPENDENCY OF CANCER CELLS: A NEW THERAPEUTIC APPROACH?.BULLETIN DU CANCER. VOL. 95. ISSUE 1. P. 69-76 | 50 | 86% | 6 |
| 7 | CELLARIER, E , DURANDO, X , VASSON, MP , FARGES, MC , DEMIDEN, A , MAURIZIS, JC , MADELMONT, JC , CHOLLET, P , (2003) METHIONINE DEPENDENCY AND CANCER TREATMENT.CANCER TREATMENT REVIEWS. VOL. 29. ISSUE 6. P. 489-499 | 62 | 70% | 74 |
| 8 | TAN, YY , XU, MX , HOFFMAN, RM , (2010) BROAD SELECTIVE EFFICACY OF RECOMBINANT METHIONINASE AND POLYETHYLENE GLYCOL-MODIFIED RECOMBINANT METHIONINASE ON CANCER CELLS IN VITRO.ANTICANCER RESEARCH. VOL. 30. ISSUE 4. P. 1041-1046 | 41 | 87% | 12 |
| 9 | SCHRAMM, VL , (2013) TRANSITION STATES, ANALOGUES, AND DRUG DEVELOPMENT.ACS CHEMICAL BIOLOGY. VOL. 8. ISSUE 1. P. 71-81 | 45 | 61% | 36 |
| 10 | BERTINO, JR , WAUD, WR , PARKER, WB , LUBIN, M , (2011) TARGETING TUMORS THAT LACK METHYLTHIOADENOSINE PHOSPHORYLASE (MTAP) ACTIVITY CURRENT STRATEGIES.CANCER BIOLOGY & THERAPY. VOL. 11. ISSUE 7. P. 627-632 | 37 | 82% | 31 |
Classes with closest relation at Level 2 |