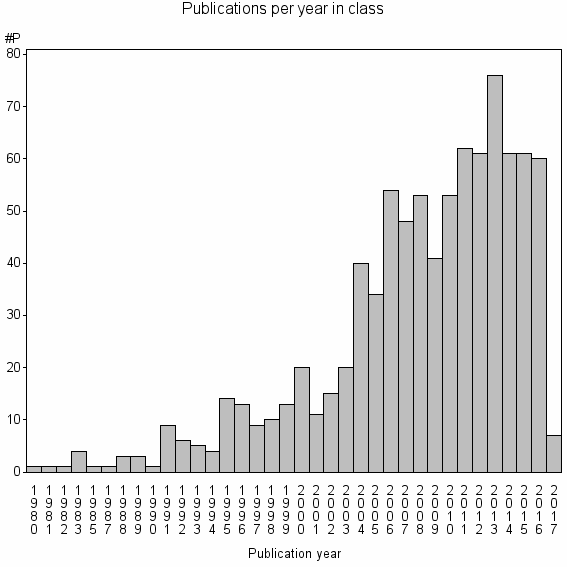
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12865 | 876 | 37.9 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 478 | 3 | PHAGE DISPLAY//CELL FREE PROTEIN SYNTHESIS//ANTIBODY ENGINEERING | 23687 |
| 1537 | 2 | CELL FREE PROTEIN SYNTHESIS//INTEIN//PROTEIN SPLICING | 7453 |
| 12865 | 1 | CELL FREE PROTEIN SYNTHESIS//CELL FREE EXPRESSION//CELL FREE | 876 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CELL FREE PROTEIN SYNTHESIS | authKW | 4678997 | 23% | 67% | 199 |
| 2 | CELL FREE EXPRESSION | authKW | 540767 | 4% | 48% | 32 |
| 3 | CELL FREE | authKW | 429844 | 4% | 33% | 37 |
| 4 | CELL FREE SYNTHETIC BIOLOGY | authKW | 351465 | 1% | 92% | 11 |
| 5 | IN VITRO SYNTHETIC BIOLOGY | authKW | 351465 | 1% | 92% | 11 |
| 6 | S30 EXTRACT | authKW | 351465 | 1% | 92% | 11 |
| 7 | CELL FREE BIOLOGY | authKW | 316873 | 1% | 91% | 10 |
| 8 | ICTAS | address | 242021 | 3% | 28% | 25 |
| 9 | IN VITRO PROTEIN SYNTHESIS | authKW | 212437 | 2% | 38% | 16 |
| 10 | CELL FREE PROTEIN EXPRESSION | authKW | 210368 | 1% | 46% | 13 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 9197 | 43% | 0% | 380 |
| 2 | Biochemical Research Methods | 3789 | 22% | 0% | 196 |
| 3 | Biochemistry & Molecular Biology | 2319 | 44% | 0% | 383 |
| 4 | Biophysics | 360 | 8% | 0% | 73 |
| 5 | Food Science & Technology | 288 | 8% | 0% | 67 |
| 6 | Cell Biology | 48 | 6% | 0% | 52 |
| 7 | Spectroscopy | 44 | 3% | 0% | 23 |
| 8 | Engineering, Chemical | 37 | 4% | 0% | 37 |
| 9 | Chemistry, Analytical | 22 | 4% | 0% | 33 |
| 10 | Chemistry, Multidisciplinary | 15 | 6% | 0% | 53 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ICTAS | 242021 | 3% | 28% | 25 |
| 2 | FINE CHEM ENGN CHEM | 145183 | 3% | 14% | 30 |
| 3 | CELL FREE SCI TECHNOL | 121584 | 3% | 12% | 29 |
| 4 | CELL FREE TECHNOL PLICAT | 104569 | 0% | 100% | 3 |
| 5 | BIOL AGR SCI | 99120 | 2% | 18% | 16 |
| 6 | BRANCH POTSDAM GOLM | 85389 | 1% | 35% | 7 |
| 7 | SIMPSON QUERREY | 83648 | 1% | 40% | 6 |
| 8 | BIOMOL MAGNET ONANCE | 81805 | 4% | 7% | 32 |
| 9 | ADV BIOTECH | 69712 | 0% | 100% | 2 |
| 10 | BRANCH BIOANALYT BIOPROC POTSDAM GOLM | 69712 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ACS SYNTHETIC BIOLOGY | 27557 | 2% | 4% | 21 |
| 2 | PROTEIN EXPRESSION AND PURIFICATION | 9191 | 4% | 1% | 35 |
| 3 | NEW BIOTECHNOLOGY | 7829 | 1% | 2% | 13 |
| 4 | BIOTECHNOLOGY AND BIOENGINEERING | 6255 | 5% | 0% | 46 |
| 5 | JOURNAL OF BIOTECHNOLOGY | 6063 | 4% | 1% | 34 |
| 6 | JOURNAL OF BIOMOLECULAR NMR | 5208 | 2% | 1% | 20 |
| 7 | BIOTECHNOLOGY PROGRESS | 5050 | 3% | 1% | 26 |
| 8 | JOURNAL OF BIOSCIENCE AND BIOENGINEERING | 4888 | 3% | 1% | 24 |
| 9 | METABOLIC ENGINEERING | 3946 | 1% | 1% | 11 |
| 10 | BIOTECHNOLOGY AND BIOPROCESS ENGINEERING | 3024 | 1% | 1% | 12 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CELL FREE PROTEIN SYNTHESIS | 4678997 | 23% | 67% | 199 | Search CELL+FREE+PROTEIN+SYNTHESIS | Search CELL+FREE+PROTEIN+SYNTHESIS |
| 2 | CELL FREE EXPRESSION | 540767 | 4% | 48% | 32 | Search CELL+FREE+EXPRESSION | Search CELL+FREE+EXPRESSION |
| 3 | CELL FREE | 429844 | 4% | 33% | 37 | Search CELL+FREE | Search CELL+FREE |
| 4 | CELL FREE SYNTHETIC BIOLOGY | 351465 | 1% | 92% | 11 | Search CELL+FREE+SYNTHETIC+BIOLOGY | Search CELL+FREE+SYNTHETIC+BIOLOGY |
| 5 | IN VITRO SYNTHETIC BIOLOGY | 351465 | 1% | 92% | 11 | Search IN+VITRO+SYNTHETIC+BIOLOGY | Search IN+VITRO+SYNTHETIC+BIOLOGY |
| 6 | S30 EXTRACT | 351465 | 1% | 92% | 11 | Search S30+EXTRACT | Search S30+EXTRACT |
| 7 | CELL FREE BIOLOGY | 316873 | 1% | 91% | 10 | Search CELL+FREE+BIOLOGY | Search CELL+FREE+BIOLOGY |
| 8 | IN VITRO PROTEIN SYNTHESIS | 212437 | 2% | 38% | 16 | Search IN+VITRO+PROTEIN+SYNTHESIS | Search IN+VITRO+PROTEIN+SYNTHESIS |
| 9 | CELL FREE PROTEIN EXPRESSION | 210368 | 1% | 46% | 13 | Search CELL+FREE+PROTEIN+EXPRESSION | Search CELL+FREE+PROTEIN+EXPRESSION |
| 10 | IN VITRO METABOLIC ENGINEERING | 209137 | 1% | 100% | 6 | Search IN+VITRO+METABOLIC+ENGINEERING | Search IN+VITRO+METABOLIC+ENGINEERING |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | PEREZ, JG , STARK, JC , JEWETT, MC , (2016) CELL-FREE SYNTHETIC BIOLOGY: ENGINEERING BEYOND THE CELL.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 8. ISSUE 12. P. - | 97 | 57% | 0 |
| 2 | BROADBENT, A , BRADLEY, WT , BUNDY, BC , SCHINN, SM , (2016) PROTEIN SYNTHESIS DIRECTLY FROM PCR: PROGRESS AND APPLICATIONS OF CELL-FREE PROTEIN SYNTHESIS WITH LINEAR DNA.NEW BIOTECHNOLOGY. VOL. 33. ISSUE 4. P. 480 -487 | 67 | 72% | 0 |
| 3 | CARLSON, ED , GAN, R , HODGMAN, CE , JEWETT, MC , (2012) CELL-FREE PROTEIN SYNTHESIS: APPLICATIONS COME OF AGE.BIOTECHNOLOGY ADVANCES. VOL. 30. ISSUE 5. P. 1185-1194 | 55 | 64% | 102 |
| 4 | KWON, YC , JEWETT, MC , (2015) HIGH-THROUGHPUT PREPARATION METHODS OF CRUDE EXTRACT FOR ROBUST CELL-FREE PROTEIN SYNTHESIS.SCIENTIFIC REPORTS. VOL. 5. ISSUE . P. - | 41 | 91% | 7 |
| 5 | HARBERS, M , (2014) WHEAT GERM SYSTEMS FOR CELL-FREE PROTEIN EXPRESSION.FEBS LETTERS. VOL. 588. ISSUE 17. P. 2762 -2773 | 62 | 55% | 13 |
| 6 | ZEMELLA, A , THORING, L , HOFFMEISTER, C , KUBICK, S , (2015) CELL-FREE PROTEIN SYNTHESIS: PROS AND CONS OF PROKARYOTIC AND EUKARYOTIC SYSTEMS.CHEMBIOCHEM. VOL. 16. ISSUE 17. P. 2420 -2431 | 67 | 49% | 6 |
| 7 | SCHWARZ, D , DOTSCH, V , BERNHARD, F , (2008) PRODUCTION OF MEMBRANE PROTEINS USING CELL-FREE EXPRESSION SYSTEMS.PROTEOMICS. VOL. 8. ISSUE 19. P. 3933-3946 | 49 | 75% | 57 |
| 8 | HENRICH, E , HEIN, C , DOTSCH, V , BERNHARD, F , (2015) MEMBRANE PROTEIN PRODUCTION IN ESCHERICHIA COLI CELL-FREE LYSATES.FEBS LETTERS. VOL. 589. ISSUE 15. P. 1713 -1722 | 59 | 52% | 9 |
| 9 | ANDERSON, MJ , STARK, JC , HODGMAN, CE , JEWETT, MC , (2015) ENERGIZING EUKARYOTIC CELL-FREE PROTEIN SYNTHESIS WITH GLUCOSE METABOLISM.FEBS LETTERS. VOL. 589. ISSUE 15. P. 1723 -1727 | 33 | 97% | 1 |
| 10 | CATHERINE, C , LEE, KH , OH, SJ , KIM, DM , (2013) CELL-FREE PLATFORMS FOR FLEXIBLE EXPRESSION AND SCREENING OF ENZYMES.BIOTECHNOLOGY ADVANCES. VOL. 31. ISSUE 6. P. 797-803 | 41 | 76% | 10 |
Classes with closest relation at Level 1 |