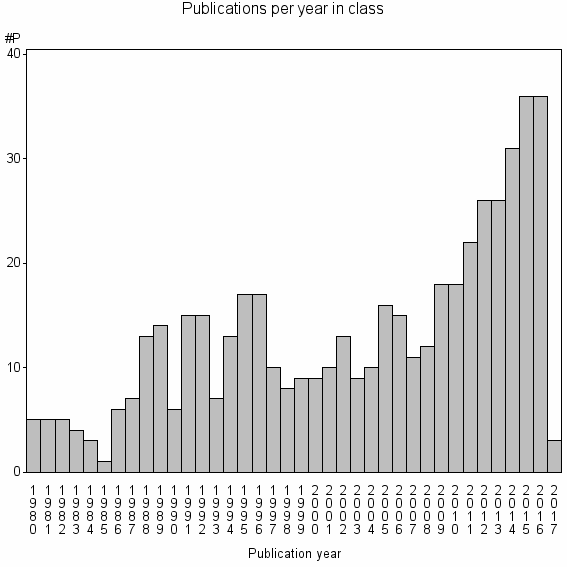
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 19522 | 501 | 48.8 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 142 | 3 | DNA REPAIR//INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH | 64602 |
| 160 | 2 | WERNER SYNDROME//DNA POLYMERASE//DNA REPAIR | 22798 |
| 19522 | 1 | REPLICATION FORK BARRIER//FORK ARREST//REPLICATION TERMINATOR PROTEIN | 501 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | REPLICATION FORK BARRIER | authKW | 507895 | 2% | 83% | 10 |
| 2 | FORK ARREST | authKW | 373303 | 1% | 88% | 7 |
| 3 | REPLICATION TERMINATOR PROTEIN | authKW | 365687 | 1% | 100% | 6 |
| 4 | R LOOPS | authKW | 361977 | 3% | 42% | 14 |
| 5 | REPLICATION TERMINATION | authKW | 352620 | 2% | 64% | 9 |
| 6 | TERMINATION OF REPLICATION | authKW | 331824 | 1% | 78% | 7 |
| 7 | ANDALUZ BIOL MOL MED REGENERAT CABIMER | address | 267871 | 4% | 22% | 20 |
| 8 | TERMINATOR PROTEIN | authKW | 243791 | 1% | 100% | 4 |
| 9 | REPLICATION TERMINUS | authKW | 217668 | 1% | 71% | 5 |
| 10 | DNA TERMINATOR | authKW | 182843 | 1% | 100% | 3 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 3223 | 65% | 0% | 326 |
| 2 | Cell Biology | 2266 | 35% | 0% | 175 |
| 3 | Genetics & Heredity | 1347 | 22% | 0% | 112 |
| 4 | Microbiology | 585 | 15% | 0% | 77 |
| 5 | Developmental Biology | 183 | 4% | 0% | 21 |
| 6 | Biophysics | 27 | 4% | 0% | 19 |
| 7 | Toxicology | 22 | 3% | 0% | 14 |
| 8 | Biology | 10 | 2% | 0% | 10 |
| 9 | Biochemical Research Methods | 3 | 2% | 0% | 9 |
| 10 | Mycology | 2 | 0% | 0% | 2 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ANDALUZ BIOL MOL MED REGENERAT CABIMER | 267871 | 4% | 22% | 20 |
| 2 | BIOL CELULAR DESARROLLO | 160098 | 3% | 15% | 17 |
| 3 | DSBB | 121888 | 1% | 33% | 6 |
| 4 | ANDALUZ BIOL MOL MED REGENERAT CAMBIMER | 60948 | 0% | 100% | 1 |
| 5 | BIOL CELULAR DESSARROLLO | 60948 | 0% | 100% | 1 |
| 6 | BIOL CELULOR | 60948 | 0% | 100% | 1 |
| 7 | BIONANOSC | 60948 | 0% | 100% | 1 |
| 8 | CLIN INVEST 808 | 60948 | 0% | 100% | 1 |
| 9 | CNRSUMR338 | 60948 | 0% | 100% | 1 |
| 10 | ECCH120 | 60948 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR CELL | 4446 | 4% | 0% | 20 |
| 2 | MOLECULAR MICROBIOLOGY | 3440 | 5% | 0% | 26 |
| 3 | CELL | 2944 | 5% | 0% | 27 |
| 4 | PLOS GENETICS | 2903 | 3% | 0% | 17 |
| 5 | GENES & DEVELOPMENT | 2631 | 4% | 0% | 18 |
| 6 | DNA REPAIR | 2413 | 2% | 0% | 9 |
| 7 | NUCLEIC ACIDS RESEARCH | 2306 | 8% | 0% | 39 |
| 8 | GENES | 1291 | 1% | 1% | 3 |
| 9 | JOURNAL OF MOLECULAR BIOLOGY | 1236 | 4% | 0% | 22 |
| 10 | EMBO JOURNAL | 1130 | 4% | 0% | 18 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | REPLICATION FORK BARRIER | 507895 | 2% | 83% | 10 | Search REPLICATION+FORK+BARRIER | Search REPLICATION+FORK+BARRIER |
| 2 | FORK ARREST | 373303 | 1% | 88% | 7 | Search FORK+ARREST | Search FORK+ARREST |
| 3 | REPLICATION TERMINATOR PROTEIN | 365687 | 1% | 100% | 6 | Search REPLICATION+TERMINATOR+PROTEIN | Search REPLICATION+TERMINATOR+PROTEIN |
| 4 | R LOOPS | 361977 | 3% | 42% | 14 | Search R+LOOPS | Search R+LOOPS |
| 5 | REPLICATION TERMINATION | 352620 | 2% | 64% | 9 | Search REPLICATION+TERMINATION | Search REPLICATION+TERMINATION |
| 6 | TERMINATION OF REPLICATION | 331824 | 1% | 78% | 7 | Search TERMINATION+OF+REPLICATION | Search TERMINATION+OF+REPLICATION |
| 7 | TERMINATOR PROTEIN | 243791 | 1% | 100% | 4 | Search TERMINATOR+PROTEIN | Search TERMINATOR+PROTEIN |
| 8 | REPLICATION TERMINUS | 217668 | 1% | 71% | 5 | Search REPLICATION+TERMINUS | Search REPLICATION+TERMINUS |
| 9 | DNA TERMINATOR | 182843 | 1% | 100% | 3 | Search DNA+TERMINATOR | Search DNA+TERMINATOR |
| 10 | 2D AGAROSE GEL ELECTROPHORESIS | 162525 | 1% | 67% | 4 | Search 2D+AGAROSE+GEL+ELECTROPHORESIS | Search 2D+AGAROSE+GEL+ELECTROPHORESIS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | BASTIA, D , ZAMAN, S , (2014) MECHANISM AND PHYSIOLOGICAL SIGNIFICANCE OF PROGRAMMED REPLICATION TERMINATION.SEMINARS IN CELL & DEVELOPMENTAL BIOLOGY. VOL. 30. ISSUE . P. 165 -173 | 60 | 69% | 5 |
| 2 | GAILLARD, H , AGUILERA, A , (2016) TRANSCRIPTION AS A THREAT TO GENOME INTEGRITY.ANNUAL REVIEW OF BIOCHEMISTRY, VOL 85. VOL. 85. ISSUE . P. 291 -317 | 71 | 41% | 2 |
| 3 | GARCIA-MUSE, T , AGUILERA, A , (2016) TRANSCRIPTION-REPLICATION CONFLICTS: HOW THEY OCCUR AND HOW THEY ARE RESOLVED.NATURE REVIEWS MOLECULAR CELL BIOLOGY. VOL. 17. ISSUE 9. P. 553 -563 | 49 | 49% | 3 |
| 4 | NEYLON, C , KRALICEK, AV , HILL, TM , DIXON, NE , (2005) REPLICATION TERMINATION IN ESCHERICHIA COLI: STRUCTURE AND ANTIHELICASE ACTIVITY OF THE TUS-TER COMPLEX.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 69. ISSUE 3. P. 501-+ | 73 | 46% | 62 |
| 5 | AGUILERA, A , GAILLARD, H , (2014) TRANSCRIPTION AND RECOMBINATION: WHEN RNA MEETS DNA.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 6. ISSUE 8. P. - | 66 | 42% | 7 |
| 6 | HAMPERL, S , CIMPRICH, KA , (2016) CONFLICT RESOLUTION IN THE GENOME: HOW TRANSCRIPTION AND REPLICATION MAKE IT WORK.CELL. VOL. 167. ISSUE 6. P. 1455 -1467 | 50 | 52% | 0 |
| 7 | SANTOS-PEREIRA, JM , AGUILERA, A , (2015) R LOOPS: NEW MODULATORS OF GENOME DYNAMICS AND FUNCTION.NATURE REVIEWS GENETICS. VOL. 16. ISSUE 10. P. 583 -597 | 48 | 38% | 29 |
| 8 | MIRKIN, EV , MIRKIN, SM , (2007) REPLICATION FORK STALLING AT NATURAL IMPEDIMENTS.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 71. ISSUE 1. P. 13-35 | 80 | 28% | 196 |
| 9 | GAILLARD, H , HERRERA-MOYANO, E , AGUILERA, A , (2013) TRANSCRIPTION-ASSOCIATED GENOME INSTABILITY.CHEMICAL REVIEWS. VOL. 113. ISSUE 11. P. 8638-8661 | 88 | 28% | 11 |
| 10 | LABIB, K , HODGSON, B , (2007) REPLICATION FORK BARRIERS: PAUSING FOR A BREAK OR STALLING FOR TIME?.EMBO REPORTS. VOL. 8. ISSUE 4. P. 346-353 | 44 | 55% | 75 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 16463 | MRNA EXPORT//NXF1//THO COMPLEX |
| 2 | 9979 | DNA SEGREGATION//FTSK//PARB |
| 3 | 13276 | UVRB//UVRD//UVRABC |
| 4 | 8862 | ORIC//DNAA//DNAA PROTEIN |
| 5 | 12602 | HOLLIDAY JUNCTION//BRANCH MIGRATION//RUVAB |
| 6 | 16834 | SECT GENE STRUCT DIS//REPEAT INSTABILITY//MUTS BETA |
| 7 | 4671 | RAD51//RAD52//HOMOLOGOUS RECOMBINATION |
| 8 | 4863 | CLAMP LOADER//REPLICATION FACTOR C//PRIMOSOME |
| 9 | 7715 | RNA POLYMERASE I//MOL BIOL CELL 2//UBF |
| 10 | 577 | DNA REPLICATION//GEMININ//CDC6 |