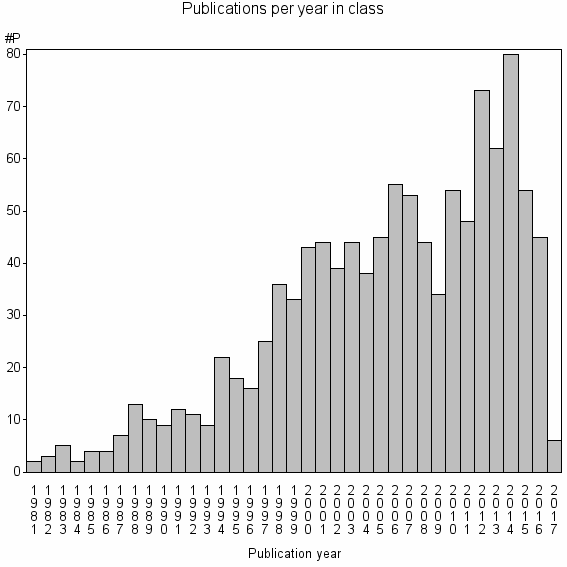
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 9979 | 1102 | 47.2 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 33 | 3 | JOURNAL OF BACTERIOLOGY//MICROBIOLOGY//MOLECULAR MICROBIOLOGY | 107033 |
| 371 | 2 | RNA POLYMERASE//HFQ//JOURNAL OF BACTERIOLOGY | 17271 |
| 9979 | 1 | DNA SEGREGATION//FTSK//PARB | 1102 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DNA SEGREGATION | authKW | 481017 | 2% | 69% | 25 |
| 2 | FTSK | authKW | 369419 | 2% | 67% | 20 |
| 3 | PARB | authKW | 362042 | 1% | 93% | 14 |
| 4 | PLASMID PARTITIONING | authKW | 270316 | 2% | 51% | 19 |
| 5 | PARAB | authKW | 221660 | 1% | 100% | 8 |
| 6 | BACTERIAL CHROMOSOME SEGREGATION | authKW | 197905 | 1% | 71% | 10 |
| 7 | DIMER RESOLUTION | authKW | 166245 | 1% | 100% | 6 |
| 8 | CHROMOSOME SEGREGATION | authKW | 157969 | 5% | 11% | 52 |
| 9 | MICROBIOL GENET MOL | address | 155922 | 5% | 11% | 50 |
| 10 | SPO0J | authKW | 150849 | 1% | 78% | 7 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Microbiology | 13250 | 46% | 0% | 504 |
| 2 | Biochemistry & Molecular Biology | 5314 | 57% | 0% | 629 |
| 3 | Cell Biology | 2012 | 23% | 0% | 254 |
| 4 | Genetics & Heredity | 948 | 13% | 0% | 147 |
| 5 | Developmental Biology | 163 | 3% | 0% | 31 |
| 6 | Biophysics | 146 | 5% | 0% | 58 |
| 7 | Mathematical & Computational Biology | 42 | 1% | 0% | 16 |
| 8 | Biology | 36 | 2% | 0% | 26 |
| 9 | Biotechnology & Applied Microbiology | 24 | 4% | 0% | 39 |
| 10 | Biochemical Research Methods | 16 | 2% | 0% | 25 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MICROBIOL GENET MOL | 155922 | 5% | 11% | 50 |
| 2 | THEORET BIOL UNIT | 98952 | 0% | 71% | 5 |
| 3 | GENE REGULAT CHROMOSOME BIOL | 89725 | 2% | 12% | 27 |
| 4 | TRANSCRIPT CONTROL SECT | 75417 | 1% | 39% | 7 |
| 5 | MOL GENET CELL OBSERV | 55411 | 0% | 50% | 4 |
| 6 | SECT THEORY COMPLEX FLUIDS | 49471 | 0% | 36% | 5 |
| 7 | STANFORD PROT NUCLE ACID IL | 41558 | 0% | 50% | 3 |
| 8 | BIOCHEM MICROBIOL UNIT | 31167 | 0% | 38% | 3 |
| 9 | UMR MICROORGANISM GENOM 7238 | 31167 | 0% | 38% | 3 |
| 10 | GENOM PHYS GRP | 30109 | 0% | 22% | 5 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR MICROBIOLOGY | 54665 | 14% | 1% | 153 |
| 2 | PLASMID | 15774 | 3% | 2% | 34 |
| 3 | JOURNAL OF BACTERIOLOGY | 15168 | 12% | 0% | 135 |
| 4 | CURRENT OPINION IN MICROBIOLOGY | 4883 | 2% | 1% | 18 |
| 5 | EMBO JOURNAL | 3370 | 4% | 0% | 46 |
| 6 | JOURNAL OF STRUCTURAL BIOLOGY | 3169 | 2% | 1% | 20 |
| 7 | JOURNAL OF MOLECULAR MICROBIOLOGY AND BIOTECHNOLOGY | 2193 | 1% | 1% | 8 |
| 8 | PLOS GENETICS | 2192 | 2% | 0% | 22 |
| 9 | GENES & DEVELOPMENT | 2108 | 2% | 0% | 24 |
| 10 | JOURNAL OF MOLECULAR BIOLOGY | 1661 | 3% | 0% | 38 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA SEGREGATION | 481017 | 2% | 69% | 25 | Search DNA+SEGREGATION | Search DNA+SEGREGATION |
| 2 | FTSK | 369419 | 2% | 67% | 20 | Search FTSK | Search FTSK |
| 3 | PARB | 362042 | 1% | 93% | 14 | Search PARB | Search PARB |
| 4 | PLASMID PARTITIONING | 270316 | 2% | 51% | 19 | Search PLASMID+PARTITIONING | Search PLASMID+PARTITIONING |
| 5 | PARAB | 221660 | 1% | 100% | 8 | Search PARAB | Search PARAB |
| 6 | BACTERIAL CHROMOSOME SEGREGATION | 197905 | 1% | 71% | 10 | Search BACTERIAL+CHROMOSOME+SEGREGATION | Search BACTERIAL+CHROMOSOME+SEGREGATION |
| 7 | DIMER RESOLUTION | 166245 | 1% | 100% | 6 | Search DIMER+RESOLUTION | Search DIMER+RESOLUTION |
| 8 | CHROMOSOME SEGREGATION | 157969 | 5% | 11% | 52 | Search CHROMOSOME+SEGREGATION | Search CHROMOSOME+SEGREGATION |
| 9 | SPO0J | 150849 | 1% | 78% | 7 | Search SPO0J | Search SPO0J |
| 10 | NUCLEOID | 131114 | 3% | 15% | 31 | Search NUCLEOID | Search NUCLEOID |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GERDES, K , HOWARD, M , SZARDENINGS, F , (2010) PUSHING AND PULLING IN PROKARYOTIC DNA SEGREGATION.CELL. VOL. 141. ISSUE 6. P. 927-942 | 108 | 72% | 149 |
| 2 | MIERZEJEWSKA, J , JAGURA-BURDZY, G , (2012) PROKARYOTIC PARA-PARB-PARS SYSTEM LINKS BACTERIAL CHROMOSOME SEGREGATION WITH THE CELL CYCLE.PLASMID. VOL. 67. ISSUE 1. P. 1-14 | 116 | 78% | 25 |
| 3 | HAYES, F , BARILLA, D , (2006) THE BACTERIAL SEGROSOME: A DYNAMIC NUCLEOPROTEIN MACHINE FOR DNA TRAFFICKING AND SEGREGATION.NATURE REVIEWS MICROBIOLOGY. VOL. 4. ISSUE 2. P. 133-143 | 106 | 84% | 96 |
| 4 | BOUET, JY , STOUF, M , LEBAILLY, E , CORNET, F , (2014) MECHANISMS FOR CHROMOSOME SEGREGATION.CURRENT OPINION IN MICROBIOLOGY. VOL. 22. ISSUE . P. 60 -65 | 81 | 100% | 6 |
| 5 | WANG, XD , LLOPIS, PM , RUDNER, DZ , (2013) ORGANIZATION AND SEGREGATION OF BACTERIAL CHROMOSOMES.NATURE REVIEWS GENETICS. VOL. 14. ISSUE 3. P. 191-203 | 91 | 65% | 78 |
| 6 | SCHUMACHER, MA , (2008) STRUCTURAL BIOLOGY OF PLASMID PARTITION: UNCOVERING THE MOLECULAR MECHANISMS OF DNA SEGREGATION.BIOCHEMICAL JOURNAL. VOL. 412. ISSUE . P. 1-18 | 106 | 81% | 33 |
| 7 | EBERSBACH, G , GERDES, K , (2005) PLASMID SEGREGATION MECHANISMS.ANNUAL REVIEW OF GENETICS. VOL. 39. ISSUE . P. 453 -479 | 106 | 75% | 147 |
| 8 | SURTEES, JA , FUNNELL, BE , (2003) PLASMID AND CHROMOSOME TRAFFIC CONTROL: HOW PARA AND PARB DRIVE PARTITION.CURRENT TOPICS IN DEVELOPMENTAL BIOLOGY, VOL 56. VOL. 56. ISSUE . P. 145-180 | 112 | 85% | 34 |
| 9 | SALJE, J , (2010) PLASMID SEGREGATION: HOW TO SURVIVE AS AN EXTRA PIECE OF DNA.CRITICAL REVIEWS IN BIOCHEMISTRY AND MOLECULAR BIOLOGY. VOL. 45. ISSUE 4. P. 296 -317 | 108 | 69% | 20 |
| 10 | BADRINARAYANAN, A , LE, TBK , LAUB, MT , (2015) BACTERIAL CHROMOSOME ORGANIZATION AND SEGREGATION.ANNUAL REVIEW OF CELL AND DEVELOPMENTAL BIOLOGY, VOL 31. VOL. 31. ISSUE . P. 171 -199 | 102 | 60% | 7 |
Classes with closest relation at Level 1 |