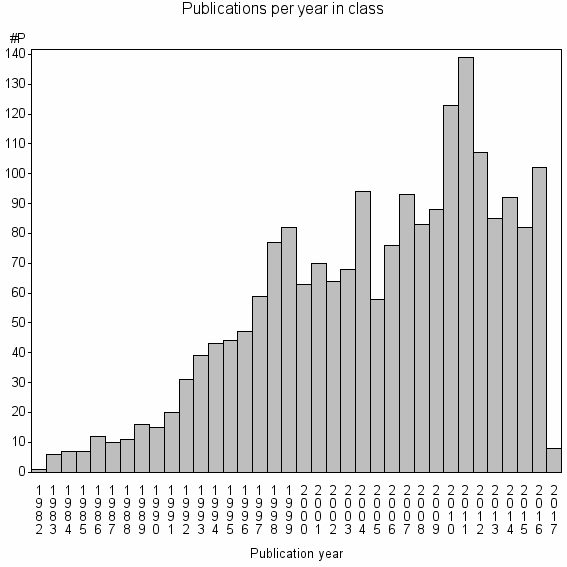
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 3273 | 2022 | 51.5 | 91% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 71 | 3 | GENETICS & HEREDITY//CHROMATIN//CELL BIOLOGY | 83302 |
| 343 | 2 | EZH2//CHROMATIN//POLYCOMB | 17754 |
| 3273 | 1 | NUCLEOSOME POSITIONING//ISWI//NUCLEOSOME | 2022 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NUCLEOSOME POSITIONING | authKW | 870527 | 5% | 56% | 103 |
| 2 | ISWI | authKW | 537968 | 2% | 76% | 47 |
| 3 | NUCLEOSOME | authKW | 399641 | 8% | 16% | 167 |
| 4 | SWI SNF | authKW | 262385 | 3% | 26% | 68 |
| 5 | CHROMATIN REMODELING | authKW | 201556 | 5% | 12% | 108 |
| 6 | CHROMATIN | authKW | 131421 | 10% | 4% | 212 |
| 7 | NURF | authKW | 121799 | 1% | 73% | 11 |
| 8 | ATP DEPENDENT CHROMATIN REMODELING | authKW | 102062 | 1% | 52% | 13 |
| 9 | CHROMATIN GENE EXP S SECT | address | 97053 | 1% | 43% | 15 |
| 10 | MNASE SEQ | authKW | 96635 | 0% | 80% | 8 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 16022 | 72% | 0% | 1448 |
| 2 | Cell Biology | 10878 | 38% | 0% | 766 |
| 3 | Genetics & Heredity | 3057 | 17% | 0% | 347 |
| 4 | Biophysics | 1221 | 10% | 0% | 199 |
| 5 | Biochemical Research Methods | 709 | 7% | 0% | 144 |
| 6 | Mathematical & Computational Biology | 546 | 3% | 0% | 67 |
| 7 | Biotechnology & Applied Microbiology | 485 | 8% | 0% | 161 |
| 8 | Developmental Biology | 465 | 3% | 0% | 69 |
| 9 | Biology | 86 | 3% | 0% | 52 |
| 10 | Mycology | 29 | 1% | 0% | 13 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CHROMATIN GENE EXP S SECT | 97053 | 1% | 43% | 15 |
| 2 | GENOME ERS | 62197 | 1% | 16% | 26 |
| 3 | CAGT PL GENOTYPING | 45299 | 0% | 100% | 3 |
| 4 | ADOLF BUTENANDT | 43991 | 2% | 7% | 41 |
| 5 | FDN IST PASTEUR | 40255 | 0% | 33% | 8 |
| 6 | EUKARYOT GENE REGULAT | 36794 | 1% | 12% | 20 |
| 7 | RECEPTOR BIOL GENE EXP S | 36262 | 1% | 9% | 28 |
| 8 | ADOLF BUTENANDT MOL BIOL | 36232 | 0% | 40% | 6 |
| 9 | FDN PIERRE GILLES GENNES RECH | 33973 | 0% | 75% | 3 |
| 10 | STATE UTILIZAT BAYAN OBO MULTIMETALL O | 30200 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR AND CELLULAR BIOLOGY | 13555 | 7% | 1% | 142 |
| 2 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 11498 | 2% | 2% | 49 |
| 3 | MOLECULAR CELL | 9883 | 3% | 1% | 60 |
| 4 | NATURE STRUCTURAL & MOLECULAR BIOLOGY | 8503 | 2% | 2% | 35 |
| 5 | NUCLEIC ACIDS RESEARCH | 7765 | 7% | 0% | 144 |
| 6 | GENOME RESEARCH | 6820 | 1% | 2% | 25 |
| 7 | EMBO JOURNAL | 6423 | 4% | 1% | 86 |
| 8 | CELL | 4440 | 3% | 0% | 67 |
| 9 | METHODS-A COMPANION TO METHODS IN ENZYMOLOGY | 4158 | 0% | 7% | 4 |
| 10 | GENES & DEVELOPMENT | 4041 | 2% | 1% | 45 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NUCLEOSOME POSITIONING | 870527 | 5% | 56% | 103 | Search NUCLEOSOME+POSITIONING | Search NUCLEOSOME+POSITIONING |
| 2 | ISWI | 537968 | 2% | 76% | 47 | Search ISWI | Search ISWI |
| 3 | NUCLEOSOME | 399641 | 8% | 16% | 167 | Search NUCLEOSOME | Search NUCLEOSOME |
| 4 | SWI SNF | 262385 | 3% | 26% | 68 | Search SWI+SNF | Search SWI+SNF |
| 5 | CHROMATIN REMODELING | 201556 | 5% | 12% | 108 | Search CHROMATIN+REMODELING | Search CHROMATIN+REMODELING |
| 6 | CHROMATIN | 131421 | 10% | 4% | 212 | Search CHROMATIN | Search CHROMATIN |
| 7 | NURF | 121799 | 1% | 73% | 11 | Search NURF | Search NURF |
| 8 | ATP DEPENDENT CHROMATIN REMODELING | 102062 | 1% | 52% | 13 | Search ATP+DEPENDENT+CHROMATIN+REMODELING | Search ATP+DEPENDENT+CHROMATIN+REMODELING |
| 9 | MNASE SEQ | 96635 | 0% | 80% | 8 | Search MNASE+SEQ | Search MNASE+SEQ |
| 10 | NUCLEOSOME REMODELING | 94356 | 1% | 42% | 15 | Search NUCLEOSOME+REMODELING | Search NUCLEOSOME+REMODELING |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LIELEG, C , KRIETENSTEIN, N , WALKER, M , KORBER, P , (2015) NUCLEOSOME POSITIONING IN YEASTS: METHODS, MAPS, AND MECHANISMS.CHROMOSOMA. VOL. 124. ISSUE 2. P. 131 -151 | 124 | 72% | 6 |
| 2 | CLAPIER, CR , CAIRNS, BR , (2009) THE BIOLOGY OF CHROMATIN REMODELING COMPLEXES.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 78. ISSUE . P. 273-304 | 93 | 52% | 788 |
| 3 | TEIF, VB , (2016) NUCLEOSOME POSITIONING: RESOURCES AND TOOLS ONLINE.BRIEFINGS IN BIOINFORMATICS. VOL. 17. ISSUE 5. P. 745 -757 | 109 | 62% | 3 |
| 4 | MUELLER-PLANITZ, F , KLINKER, H , BECKER, PB , (2013) NUCLEOSOME SLIDING MECHANISMS: NEW TWISTS IN A LOOPED HISTORY.NATURE STRUCTURAL & MOLECULAR BIOLOGY. VOL. 20. ISSUE 9. P. 1026-1032 | 84 | 88% | 29 |
| 5 | NARLIKAR, GJ , SUNDARAMOORTHY, R , OWEN-HUGHES, T , (2013) MECHANISMS AND FUNCTIONS OF ATP-DEPENDENT CHROMATIN-REMODELING ENZYMES.CELL. VOL. 154. ISSUE 3. P. 490 -503 | 64 | 72% | 125 |
| 6 | BECKER, PB , HORZ, W , (2002) ATP-DEPENDENT NUCLEOSOMERE MODELING.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 71. ISSUE . P. 247 -273 | 102 | 61% | 486 |
| 7 | STRUHL, K , SEGAL, E , (2013) DETERMINANTS OF NUCLEOSOME POSITIONING.NATURE STRUCTURAL & MOLECULAR BIOLOGY. VOL. 20. ISSUE 3. P. 267-273 | 53 | 78% | 151 |
| 8 | JANSEN, A , VERSTREPEN, KJ , (2011) NUCLEOSOME POSITIONING IN SACCHAROMYCES CEREVISIAE.MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS. VOL. 75. ISSUE 2. P. 301-+ | 114 | 58% | 34 |
| 9 | RANDO, OJ , WINSTON, F , (2012) CHROMATIN AND TRANSCRIPTION IN YEAST.GENETICS. VOL. 190. ISSUE 2. P. 351-387 | 170 | 32% | 80 |
| 10 | ZHOU, CY , JOHNSON, SL , GAMARRA, NI , NARLIKAR, GJ , (2016) MECHANISMS OF ATP-DEPENDENT CHROMATIN REMODELING MOTORS.ANNUAL REVIEW OF BIOPHYSICS, VOL 45. VOL. 45. ISSUE . P. 153 -181 | 94 | 63% | 1 |
Classes with closest relation at Level 1 |