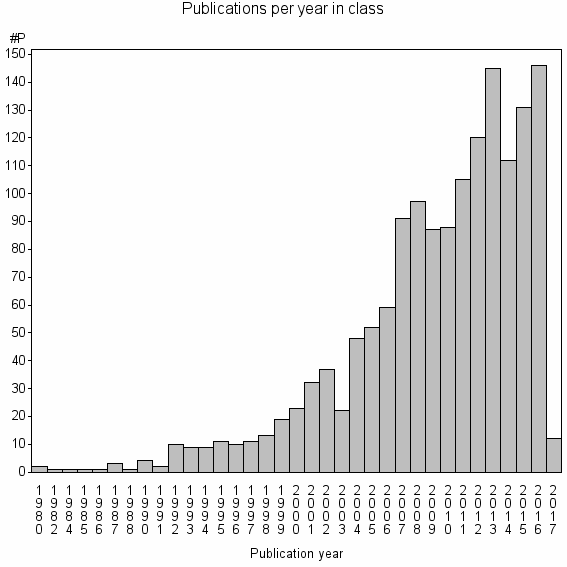
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 6198 | 1515 | 53.0 | 94% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 153 | 3 | BIOCHEMISTRY & MOLECULAR BIOLOGY//RIBOSOME//RNA | 62593 |
| 156 | 2 | NUCLEAR PORE COMPLEX//PRE MRNA SPLICING//SPLICEOSOME | 22915 |
| 6198 | 1 | P BODIES//STRESS GRANULES//DECAPPING | 1515 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | P BODIES | authKW | 1069301 | 6% | 57% | 93 |
| 2 | STRESS GRANULES | authKW | 863424 | 7% | 38% | 113 |
| 3 | DECAPPING | authKW | 835107 | 4% | 75% | 55 |
| 4 | DEADENYLATION | authKW | 619703 | 4% | 50% | 62 |
| 5 | MRNA DECAY | authKW | 618990 | 6% | 32% | 96 |
| 6 | CCR4 NOT | authKW | 424101 | 1% | 96% | 22 |
| 7 | DEADENYLASE | authKW | 400690 | 2% | 76% | 26 |
| 8 | PROCESSING BODIES | authKW | 347692 | 2% | 51% | 34 |
| 9 | CCR4 NOT COMPLEX | authKW | 287898 | 1% | 71% | 20 |
| 10 | GW182 | authKW | 286689 | 1% | 68% | 21 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Cell Biology | 11141 | 44% | 0% | 665 |
| 2 | Biochemistry & Molecular Biology | 9196 | 63% | 0% | 960 |
| 3 | Virology | 672 | 6% | 0% | 84 |
| 4 | Genetics & Heredity | 611 | 10% | 0% | 146 |
| 5 | Biophysics | 468 | 7% | 0% | 112 |
| 6 | Developmental Biology | 171 | 3% | 0% | 38 |
| 7 | Biochemical Research Methods | 82 | 3% | 0% | 52 |
| 8 | Mycology | 75 | 1% | 0% | 16 |
| 9 | Biology | 26 | 2% | 0% | 29 |
| 10 | Microbiology | 22 | 3% | 0% | 48 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CELL SIGNAL UNIT | 96723 | 1% | 40% | 12 |
| 2 | CNRS FRE 3402 | 53740 | 0% | 67% | 4 |
| 3 | HELMHOLTZ JR GRP POSTTRANSCRIPT CONTROL GENE | 40299 | 0% | 33% | 6 |
| 4 | HIV RNA TRAFFICKING 1 | 36268 | 0% | 30% | 6 |
| 5 | ERI BIOTECMED | 34042 | 0% | 24% | 7 |
| 6 | BIOL CHEM MOL PHARMACOLHEMATOL | 26870 | 0% | 67% | 2 |
| 7 | ST PATRICK GRP BASIC ONCOLHOTEL DIEU QUEBEC | 26870 | 0% | 67% | 2 |
| 8 | ABORAT PROGRAM GENOME BIOL BIOINFORMAT | 20154 | 0% | 100% | 1 |
| 9 | ANDALUZ BIOL MOL MED REGENAT CABIMER | 20154 | 0% | 100% | 1 |
| 10 | ARN CGM | 20154 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | RNA | 41139 | 5% | 3% | 72 |
| 2 | MOLECULAR CELL | 12360 | 4% | 1% | 58 |
| 3 | WILEY INTERDISCIPLINARY REVIEWS-RNA | 12327 | 1% | 4% | 15 |
| 4 | RNA-A PUBLICATION OF THE RNA SOCIETY | 9861 | 2% | 2% | 25 |
| 5 | RNA BIOLOGY | 9651 | 1% | 2% | 21 |
| 6 | NATURE STRUCTURAL & MOLECULAR BIOLOGY | 9500 | 2% | 1% | 32 |
| 7 | BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS | 6912 | 1% | 2% | 19 |
| 8 | MOLECULAR AND CELLULAR BIOLOGY | 5278 | 5% | 0% | 77 |
| 9 | MOLECULAR BIOLOGY OF THE CELL | 4011 | 3% | 0% | 41 |
| 10 | NUCLEIC ACIDS RESEARCH | 3405 | 5% | 0% | 83 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | P BODIES | 1069301 | 6% | 57% | 93 | Search P+BODIES | Search P+BODIES |
| 2 | STRESS GRANULES | 863424 | 7% | 38% | 113 | Search STRESS+GRANULES | Search STRESS+GRANULES |
| 3 | DECAPPING | 835107 | 4% | 75% | 55 | Search DECAPPING | Search DECAPPING |
| 4 | DEADENYLATION | 619703 | 4% | 50% | 62 | Search DEADENYLATION | Search DEADENYLATION |
| 5 | MRNA DECAY | 618990 | 6% | 32% | 96 | Search MRNA+DECAY | Search MRNA+DECAY |
| 6 | CCR4 NOT | 424101 | 1% | 96% | 22 | Search CCR4+NOT | Search CCR4+NOT |
| 7 | DEADENYLASE | 400690 | 2% | 76% | 26 | Search DEADENYLASE | Search DEADENYLASE |
| 8 | PROCESSING BODIES | 347692 | 2% | 51% | 34 | Search PROCESSING+BODIES | Search PROCESSING+BODIES |
| 9 | CCR4 NOT COMPLEX | 287898 | 1% | 71% | 20 | Search CCR4+NOT+COMPLEX | Search CCR4+NOT+COMPLEX |
| 10 | GW182 | 286689 | 1% | 68% | 21 | Search GW182 | Search GW182 |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | JONAS, S , IZAURRALDE, E , (2015) NON-CODING RNA TOWARDS A MOLECULAR UNDERSTANDING OF MICRORNA-MEDIATED GENE SILENCING.NATURE REVIEWS GENETICS. VOL. 16. ISSUE 7. P. 421 -433 | 85 | 75% | 108 |
| 2 | DECKER, CJ , PARKER, R , (2012) P-BODIES AND STRESS GRANULES: POSSIBLE ROLES IN THE CONTROL OF TRANSLATION AND MRNA DEGRADATION.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 4. ISSUE 9. P. - | 102 | 77% | 127 |
| 3 | FRANKS, TM , LYKKE-ANDERSEN, J , (2008) THE CONTROL OF MRNA DECAPPING AND P-BODY FORMATION.MOLECULAR CELL. VOL. 32. ISSUE 5. P. 605-615 | 99 | 82% | 205 |
| 4 | JONAS, S , IZAURRALDE, E , (2013) THE ROLE OF DISORDERED PROTEIN REGIONS IN THE ASSEMBLY OF DECAPPING COMPLEXES AND RNP GRANULES.GENES & DEVELOPMENT. VOL. 27. ISSUE 24. P. 2628-2641 | 82 | 85% | 38 |
| 5 | JAIN, S , PARKER, R , (2013) THE DISCOVERY AND ANALYSIS OF P BODIES.TEN YEARS OF PROGRESS IN GW/P BODY RESEARCH. VOL. 768. ISSUE . P. 23 -43 | 100 | 76% | 21 |
| 6 | ARRIBAS-LAYTON, M , WU, DH , LYKKE-ANDERSEN, J , SONG, HW , (2013) STRUCTURAL AND FUNCTIONAL CONTROL OF THE EUKARYOTIC MRNA DECAPPING MACHINERY.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1829. ISSUE 6-7. P. 580 -589 | 92 | 74% | 29 |
| 7 | COLLART, MA , PANASENKO, OO , (2012) THE CCR4-NOT COMPLEX.GENE. VOL. 492. ISSUE 1. P. 42 -53 | 98 | 56% | 110 |
| 8 | COLLART, MA , (2016) THE CCR4-NOT COMPLEX IS A KEY REGULATOR OF EUKARYOTIC GENE EXPRESSION.WILEY INTERDISCIPLINARY REVIEWS-RNA. VOL. 7. ISSUE 4. P. 438 -454 | 69 | 74% | 5 |
| 9 | WAHLE, E , WINKLER, GS , (2013) RNA DECAY MACHINES: DEADENYLATION BY THE CCR4-NOT AND PAN2-PAN3 COMPLEXES.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1829. ISSUE 6-7. P. 561-570 | 84 | 66% | 56 |
| 10 | LING, SHM , QAMRA, R , SONG, HW , (2011) STRUCTURAL AND FUNCTIONAL INSIGHTS INTO EUKARYOTIC MRNA DECAPPING.WILEY INTERDISCIPLINARY REVIEWS-RNA. VOL. 2. ISSUE 2. P. 193-208 | 100 | 69% | 21 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 17891 | RRP6//MRNP BIOGENESIS METAB//DIS3 |
| 2 | 10335 | NMD//UPF1//NONSENSE MEDIATED MRNA DECAY |
| 3 | 35582 | RASGAP//G3BP1//G3BP |
| 4 | 11961 | CYTOPLASMIC POLYADENYLATION//CPEB//POLYA BINDING PROTEIN |
| 5 | 3617 | HUR//AU RICH ELEMENT//TRISTETRAPROLIN |
| 6 | 24691 | BTG2//BTG1//TOB |
| 7 | 35793 | RIBOTAG//S100A10//4SU |
| 8 | 12218 | RNA HELICASE//DEAD BOX PROTEIN//DEAD BOX |
| 9 | 5071 | RNA LOCALIZATION//OSKAR//GERM PLASM |
| 10 | 3885 | EIF4E//EIF4F//CAP ANALOG |