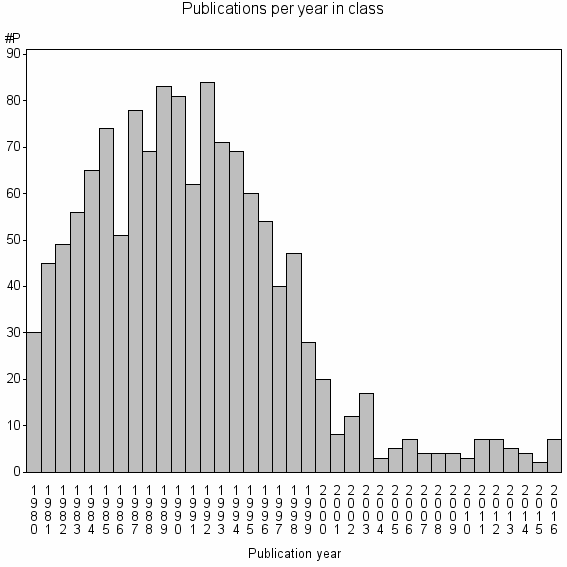
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 7907 | 1315 | 36.7 | 62% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 189 | 3 | JOURNAL OF MAGNETIC RESONANCE//MAGNETIC RESONANCE IN MEDICINE//JOURNAL OF BIOMOLECULAR NMR | 54843 |
| 866 | 2 | JOURNAL OF BIOMOLECULAR NMR//JOURNAL OF MAGNETIC RESONANCE//RESIDUAL DIPOLAR COUPLINGS | 11581 |
| 7907 | 1 | DISTANCE GEOMETRY//JOURNAL OF MAGNETIC RESONANCE SERIES B//DGGTATACC2 | 1315 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DISTANCE GEOMETRY | authKW | 103573 | 3% | 14% | 33 |
| 2 | JOURNAL OF MAGNETIC RESONANCE SERIES B | journal | 78021 | 3% | 7% | 45 |
| 3 | DGGTATACC2 | authKW | 69657 | 0% | 100% | 3 |
| 4 | SAMPLING CONFORMATION SPACE | authKW | 69657 | 0% | 100% | 3 |
| 5 | COMPLETE RELAXATION MATRIX ANALYSIS | authKW | 64493 | 0% | 56% | 5 |
| 6 | JOURNAL OF MAGNETIC RESONANCE | journal | 59809 | 12% | 2% | 152 |
| 7 | 3J COUPLING CONSTANTS | authKW | 58043 | 0% | 50% | 5 |
| 8 | TIME AVERAGED RESTRAINTS | authKW | 52242 | 0% | 75% | 3 |
| 9 | 2D NMR OF DNA | authKW | 46438 | 0% | 100% | 2 |
| 10 | COMPLETE CROSS VALIDATION | authKW | 46438 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Spectroscopy | 8771 | 23% | 0% | 298 |
| 2 | Biochemistry & Molecular Biology | 5259 | 52% | 0% | 690 |
| 3 | Physics, Atomic, Molecular & Chemical | 3195 | 20% | 0% | 266 |
| 4 | Biochemical Research Methods | 2534 | 15% | 0% | 201 |
| 5 | Biophysics | 2230 | 16% | 0% | 206 |
| 6 | Chemistry, Multidisciplinary | 581 | 16% | 0% | 212 |
| 7 | Chemistry, Physical | 106 | 9% | 0% | 122 |
| 8 | Chemistry, Organic | 17 | 3% | 0% | 45 |
| 9 | Biology | 14 | 2% | 0% | 22 |
| 10 | Computer Science, Interdisciplinary Applications | 13 | 2% | 0% | 20 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOCHEM MAGNET ONANCE IL MADISON | 23219 | 0% | 100% | 1 |
| 2 | BIOMOL NAT SCI | 23219 | 0% | 100% | 1 |
| 3 | CHEM ABT HERMANN REIN STR 3 | 23219 | 0% | 100% | 1 |
| 4 | CHIM BIOL STRUCT FONCT | 23219 | 0% | 100% | 1 |
| 5 | NMR ORGAN CHEM GRP | 23219 | 0% | 100% | 1 |
| 6 | PARIS CHIM | 23219 | 0% | 100% | 1 |
| 7 | THEOR CHEM PHYS | 23219 | 0% | 100% | 1 |
| 8 | UNITE BIOINFORMAT STURCT | 23219 | 0% | 100% | 1 |
| 9 | MOL PHYS CHEM RD UNIT | 15477 | 0% | 33% | 2 |
| 10 | BIOCHIM THEOR CNRS URA 77 | 11609 | 0% | 50% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF MAGNETIC RESONANCE SERIES B | 78021 | 3% | 7% | 45 |
| 2 | JOURNAL OF MAGNETIC RESONANCE | 59809 | 12% | 2% | 152 |
| 3 | JOURNAL OF BIOMOLECULAR NMR | 37938 | 5% | 2% | 66 |
| 4 | BIOCHEMISTRY | 8432 | 11% | 0% | 140 |
| 5 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 8021 | 3% | 1% | 33 |
| 6 | BIOPOLYMERS | 7183 | 3% | 1% | 45 |
| 7 | PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY | 6842 | 1% | 2% | 12 |
| 8 | MAGNETIC RESONANCE IN CHEMISTRY | 6713 | 3% | 1% | 41 |
| 9 | MAGNETIC RESONANCE IN BIOLOGY | 4642 | 0% | 20% | 1 |
| 10 | JOURNAL OF MAGNETIC RESONANCE SERIES A | 3814 | 1% | 1% | 13 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DISTANCE GEOMETRY | 103573 | 3% | 14% | 33 | Search DISTANCE+GEOMETRY | Search DISTANCE+GEOMETRY |
| 2 | DGGTATACC2 | 69657 | 0% | 100% | 3 | Search DGGTATACC2 | Search DGGTATACC2 |
| 3 | SAMPLING CONFORMATION SPACE | 69657 | 0% | 100% | 3 | Search SAMPLING+CONFORMATION+SPACE | Search SAMPLING+CONFORMATION+SPACE |
| 4 | COMPLETE RELAXATION MATRIX ANALYSIS | 64493 | 0% | 56% | 5 | Search COMPLETE+RELAXATION+MATRIX+ANALYSIS | Search COMPLETE+RELAXATION+MATRIX+ANALYSIS |
| 5 | 3J COUPLING CONSTANTS | 58043 | 0% | 50% | 5 | Search 3J+COUPLING+CONSTANTS | Search 3J+COUPLING+CONSTANTS |
| 6 | TIME AVERAGED RESTRAINTS | 52242 | 0% | 75% | 3 | Search TIME+AVERAGED+RESTRAINTS | Search TIME+AVERAGED+RESTRAINTS |
| 7 | 2D NMR OF DNA | 46438 | 0% | 100% | 2 | Search 2D+NMR+OF+DNA | Search 2D+NMR+OF+DNA |
| 8 | COMPLETE CROSS VALIDATION | 46438 | 0% | 100% | 2 | Search COMPLETE+CROSS+VALIDATION | Search COMPLETE+CROSS+VALIDATION |
| 9 | DG II | 46438 | 0% | 100% | 2 | Search DG+II | Search DG+II |
| 10 | DYNAMIC NMR REFINEMENT | 46438 | 0% | 100% | 2 | Search DYNAMIC+NMR+REFINEMENT | Search DYNAMIC+NMR+REFINEMENT |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VOGELI, B , (2014) THE NUCLEAR OVERHAUSER EFFECT FROM A QUANTITATIVE PERSPECTIVE.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 78. ISSUE . P. 1 -46 | 91 | 46% | 8 |
| 2 | VANDEVEN, FJM , HILBERS, CW , (1988) NUCLEIC-ACIDS AND NUCLEAR MAGNETIC-RESONANCE.EUROPEAN JOURNAL OF BIOCHEMISTRY. VOL. 178. ISSUE 1. P. 1-38 | 137 | 39% | 287 |
| 3 | LUXON, BA , GORENSTEIN, DG , (1995) COMPARISON OF X-RAY AND NMR-DETERMINED NUCLEIC ACID STRUCTURES.NUCLEAR MAGNETIC RESONANCE AND NUCLEIC ACIDS. VOL. 261. ISSUE . P. 45-73 | 64 | 75% | 17 |
| 4 | JAMES, TL , (1994) COMPUTATIONAL STRATEGIES PERTINENT TO NMR SOLUTION STRUCTURE DETERMINATION.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 4. ISSUE 2. P. 275 -284 | 58 | 73% | 19 |
| 5 | SHRIVER, JW , EDMONDSON, S , (1994) ERROR ANALYSIS OF MACROMOLECULAR STRUCTURES DETERMINED WITH NUCLEAR-MAGNETIC-RESONANCE DATA.NUMERICAL COMPUTER METHODS, PT B. VOL. 240. ISSUE . P. 415-438 | 48 | 87% | 3 |
| 6 | BRUSCHWEILER, R , CASE, DA , (1994) CHARACTERIZATION OF BIOMOLECULAR STRUCTURE AND DYNAMICS BY NMR CROSS-RELAXATION.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 26. ISSUE . P. 27 -58 | 71 | 52% | 53 |
| 7 | JAMES, TL , (1994) ASSESSMENT OF QUALITY OF DERIVED MACROMOLECULAR STRUCTURES.NUCLEAR MAGNETIC RESONANCE, PT C. VOL. 239. ISSUE . P. 416 -439 | 51 | 76% | 21 |
| 8 | GUNTERT, P , (1998) STRUCTURE CALCULATION OF BIOLOGICAL MACROMOLECULES FROM NMR DATA.QUARTERLY REVIEWS OF BIOPHYSICS. VOL. 31. ISSUE 2. P. 145 -237 | 65 | 43% | 93 |
| 9 | BRUNGER, AT , NILGES, M , (1993) COMPUTATIONAL CHALLENGES FOR MACROMOLECULAR STRUCTURE DETERMINATION BY X-RAY CRYSTALLOGRAPHY AND SOLUTION NMR-SPECTROSCOPY.QUARTERLY REVIEWS OF BIOPHYSICS. VOL. 26. ISSUE 1. P. 49 -125 | 95 | 35% | 95 |
| 10 | HOSUR, RV , GOVIL, G , MILES, HT , (1988) APPLICATION OF TWO-DIMENSIONAL NMR-SPECTROSCOPY IN THE DETERMINATION OF SOLUTION CONFORMATION OF NUCLEIC-ACIDS.MAGNETIC RESONANCE IN CHEMISTRY. VOL. 26. ISSUE 10. P. 927-944 | 58 | 73% | 54 |
Classes with closest relation at Level 1 |