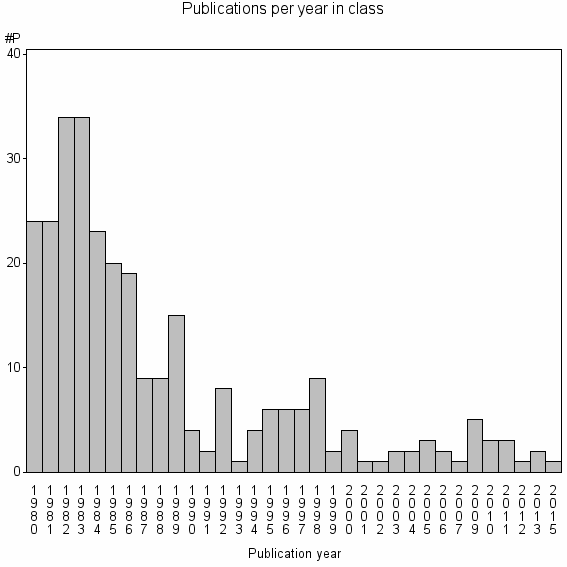
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 25121 | 290 | 31.7 | 38% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 189 | 3 | JOURNAL OF MAGNETIC RESONANCE//MAGNETIC RESONANCE IN MEDICINE//JOURNAL OF BIOMOLECULAR NMR | 54843 |
| 866 | 2 | JOURNAL OF BIOMOLECULAR NMR//JOURNAL OF MAGNETIC RESONANCE//RESIDUAL DIPOLAR COUPLINGS | 11581 |
| 25121 | 1 | KSGA//DIM1//DIMETHYLTRANSFERASE | 290 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | KSGA | authKW | 329040 | 2% | 63% | 5 |
| 2 | DIM1 | authKW | 168466 | 1% | 40% | 4 |
| 3 | DIMETHYLTRANSFERASE | authKW | 140391 | 1% | 67% | 2 |
| 4 | ANTHRANILOYL | authKW | 105294 | 0% | 100% | 1 |
| 5 | ASSIGNMENT OF IMINO PROTON SIGNALS | authKW | 105294 | 0% | 100% | 1 |
| 6 | BIOCHIM CNRS URA240 | address | 105294 | 0% | 100% | 1 |
| 7 | BIOL MCS | address | 105294 | 0% | 100% | 1 |
| 8 | DIMETHYLASE | authKW | 105294 | 0% | 100% | 1 |
| 9 | DNA SOLUTION CONFORMATION | authKW | 105294 | 0% | 100% | 1 |
| 10 | IMINO GROUPS | authKW | 105294 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 2378 | 73% | 0% | 211 |
| 2 | Biophysics | 686 | 18% | 0% | 53 |
| 3 | Biology | 27 | 3% | 0% | 10 |
| 4 | Chemistry, Multidisciplinary | 21 | 9% | 0% | 25 |
| 5 | Chemistry, Organic | 17 | 5% | 0% | 15 |
| 6 | Biochemical Research Methods | 16 | 3% | 0% | 10 |
| 7 | Spectroscopy | 4 | 2% | 0% | 5 |
| 8 | Materials Science, Textiles | 1 | 0% | 0% | 1 |
| 9 | Microbiology | 1 | 2% | 0% | 7 |
| 10 | Biotechnology & Applied Microbiology | 1 | 2% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOCHIM CNRS URA240 | 105294 | 0% | 100% | 1 |
| 2 | BIOL MCS | 105294 | 0% | 100% | 1 |
| 3 | BIOL MOL CELL BIOL ANIM BIOL | 35097 | 0% | 33% | 1 |
| 4 | STRUCT BIOL DRUG DISCOVERY | 17242 | 2% | 2% | 7 |
| 5 | EGG QUAL SAFETY UNIT | 15040 | 0% | 14% | 1 |
| 6 | MOLEC CELLULAR DEV BIOL PROGRAM | 13160 | 0% | 13% | 1 |
| 7 | STRUCT BIOL DRUG DESIGN | 11698 | 0% | 11% | 1 |
| 8 | INFORMAT EXTREMOPHILES | 6579 | 0% | 6% | 1 |
| 9 | POB 1016 | 6192 | 0% | 6% | 1 |
| 10 | STATE MOL BIOL BIOCHEM CELL BIOL | 5848 | 0% | 6% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOCHEMISTRY | 4741 | 17% | 0% | 49 |
| 2 | ANNUAL REVIEW OF BIOPHYSICS AND BIOENGINEERING | 4383 | 1% | 2% | 2 |
| 3 | NUCLEIC ACIDS RESEARCH | 4251 | 14% | 0% | 40 |
| 4 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 4052 | 4% | 0% | 11 |
| 5 | KHIMIYA PRIRODNYKH SOEDINENII | 2423 | 3% | 0% | 9 |
| 6 | JOURNAL OF CARBOHYDRATES-NUCLEOSIDES-NUCLEOTIDES | 1813 | 0% | 2% | 1 |
| 7 | POSTEPY BIOCHEMII | 1365 | 0% | 1% | 1 |
| 8 | JOURNAL OF MOLECULAR BIOLOGY | 1138 | 6% | 0% | 16 |
| 9 | CHEMIKER-ZEITUNG | 1087 | 1% | 0% | 3 |
| 10 | EUROPEAN JOURNAL OF BIOCHEMISTRY | 752 | 4% | 0% | 12 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | KSGA | 329040 | 2% | 63% | 5 | Search KSGA | Search KSGA |
| 2 | DIM1 | 168466 | 1% | 40% | 4 | Search DIM1 | Search DIM1 |
| 3 | DIMETHYLTRANSFERASE | 140391 | 1% | 67% | 2 | Search DIMETHYLTRANSFERASE | Search DIMETHYLTRANSFERASE |
| 4 | ANTHRANILOYL | 105294 | 0% | 100% | 1 | Search ANTHRANILOYL | Search ANTHRANILOYL |
| 5 | ASSIGNMENT OF IMINO PROTON SIGNALS | 105294 | 0% | 100% | 1 | Search ASSIGNMENT+OF+IMINO+PROTON+SIGNALS | Search ASSIGNMENT+OF+IMINO+PROTON+SIGNALS |
| 6 | DIMETHYLASE | 105294 | 0% | 100% | 1 | Search DIMETHYLASE | Search DIMETHYLASE |
| 7 | DNA SOLUTION CONFORMATION | 105294 | 0% | 100% | 1 | Search DNA+SOLUTION+CONFORMATION | Search DNA+SOLUTION+CONFORMATION |
| 8 | IMINO GROUPS | 105294 | 0% | 100% | 1 | Search IMINO+GROUPS | Search IMINO+GROUPS |
| 9 | IMINO PROTON RESONANCES | 105294 | 0% | 100% | 1 | Search IMINO+PROTON+RESONANCES | Search IMINO+PROTON+RESONANCES |
| 10 | ION INDUCED HYDROLYSIS | 105294 | 0% | 100% | 1 | Search ION+INDUCED+HYDROLYSIS | Search ION+INDUCED+HYDROLYSIS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HYDE, EI , (1986) IMINO PROTON NMR ASSIGNMENTS AND ION-BINDING STUDIES ON ESCHERICHIA-COLI TRANSFER RNA3GLY.EUROPEAN JOURNAL OF BIOCHEMISTRY. VOL. 155. ISSUE 1. P. 57-68 | 23 | 92% | 7 |
| 2 | HALL, KB , SAMPSON, JR , UHLENBECK, OC , REDFIELD, AG , (1989) STRUCTURE OF AN UNMODIFIED TRANSFER-RNA MOLECULE.BIOCHEMISTRY. VOL. 28. ISSUE 14. P. 5794-5801 | 13 | 87% | 117 |
| 3 | HARDIN, CC , GOLLNICK, P , HOROWITZ, J , (1988) PARTIAL ASSIGNMENT OF RESONANCES IN THE F-19 NUCLEAR MAGNETIC-RESONANCE SPECTRA OF 5-FLUOROURACIL-SUBSTITUTED TRANSFER-RNAS.BIOCHEMISTRY. VOL. 27. ISSUE 1. P. 487-495 | 15 | 94% | 7 |
| 4 | FLANAGAN, JM , JACOBSON, KB , (1988) EFFECT OF ZINC IONS ON TRANSFER-RNA STRUCTURE - IMINO PROTON NMR-SPECTROSCOPY.BIOCHEMISTRY. VOL. 27. ISSUE 15. P. 5778-5785 | 15 | 94% | 3 |
| 5 | CHOI, BS , REDFIELD, AG , (1992) NMR-STUDY OF NITROGEN-15-LABELED ESCHERICHIA-COLI VALINE TRANSFER-RNA.BIOCHEMISTRY. VOL. 31. ISSUE 51. P. 12799-12802 | 16 | 70% | 6 |
| 6 | CHOI, BS , REDFIELD, AG , (1986) NMR-STUDY OF ISOLEUCINE TRANSFER-RNA FROM THERMUS-THERMOPHILUS.BIOCHEMISTRY. VOL. 25. ISSUE 7. P. 1529-1534 | 18 | 82% | 3 |
| 7 | TU, HM , YANG, YS , MENG, JX , GONG, ML , WU, YJ , ZHANG, HJ , (1998) PROBING LANTHANIDE IONS BINDING ON TRNA(PHE) BY H-1 NMR.JOURNAL OF RARE EARTHS. VOL. 16. ISSUE 3. P. 161-166 | 10 | 83% | 0 |
| 8 | GOLLNICK, P , HARDIN, CC , HOROWITZ, J , (1987) F-19 NUCLEAR-MAGNETIC-RESONANCE AS A PROBE OF ANTICODON STRUCTURE IN 5-FLUOROURACIL-SUBSTITUTED ESCHERICHIA-COLI TRANSFER-RNA.JOURNAL OF MOLECULAR BIOLOGY. VOL. 197. ISSUE 3. P. 571-584 | 18 | 72% | 7 |
| 9 | TU, HM , WU, YJ , YANG, YS , GONG, ML , ZHANG, HJ , (1998) EFFECT OF LANTHANUM IONS ON TRNA(PHE) STRUCTURE: IMINO PROTON NMR SPECTROSCOPY.CHINESE SCIENCE BULLETIN. VOL. 43. ISSUE 6. P. 490-493 | 10 | 77% | 0 |
| 10 | STRIKER, G , LABUDA, D , VEGAMARTIN, MD , (1989) THE 3 CONFORMATIONS OF THE ANTICODON LOOP OF YEAST TRANSFER RNAPHE.JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS. VOL. 7. ISSUE 2. P. 235-255 | 14 | 64% | 13 |
Classes with closest relation at Level 1 |