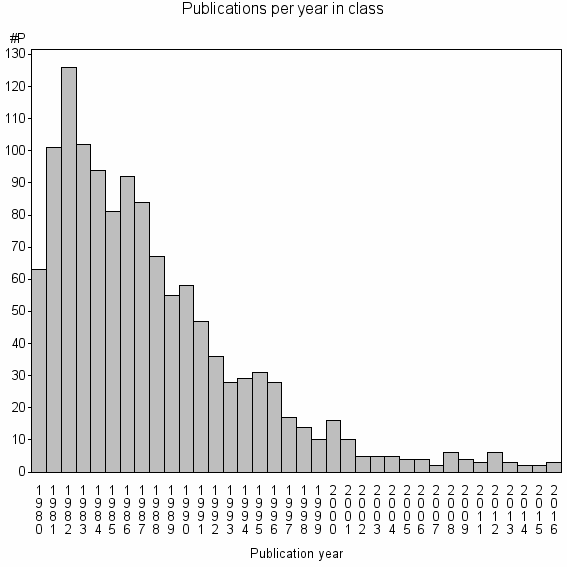
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 8539 | 1243 | 32.9 | 52% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 153 | 3 | BIOCHEMISTRY & MOLECULAR BIOLOGY//RIBOSOME//RNA | 62593 |
| 1020 | 2 | NUCLEOLUS//RIBOSOME BIOGENESIS//SNORNA | 10264 |
| 8539 | 1 | 5S RRNA//5S RIBOSOMAL RNA//EXPANSION SEGMENT | 1243 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | 5S RRNA | authKW | 114959 | 2% | 20% | 23 |
| 2 | 5S RIBOSOMAL RNA | authKW | 87707 | 1% | 24% | 15 |
| 3 | EXPANSION SEGMENT | authKW | 86498 | 1% | 39% | 9 |
| 4 | RIBOSOMAL PROTEIN L5 | authKW | 73687 | 0% | 50% | 6 |
| 5 | HIDDEN BREAK | authKW | 65502 | 0% | 67% | 4 |
| 6 | ARRHENIUS S E5 | address | 63158 | 0% | 43% | 6 |
| 7 | PHYSIOCOCHEM MED | address | 49128 | 0% | 100% | 2 |
| 8 | RIBOSOME RECONSTITUTION | authKW | 49128 | 0% | 100% | 2 |
| 9 | NUCLEIC ACIDS RESEARCH | journal | 38973 | 20% | 1% | 250 |
| 10 | COMPENSATORY SLIPPAGE | authKW | 32751 | 0% | 67% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 9822 | 72% | 0% | 889 |
| 2 | Evolutionary Biology | 1261 | 7% | 0% | 87 |
| 3 | Biophysics | 1153 | 12% | 0% | 148 |
| 4 | Genetics & Heredity | 930 | 13% | 0% | 156 |
| 5 | Microbiology | 633 | 11% | 0% | 133 |
| 6 | Biology | 397 | 6% | 0% | 71 |
| 7 | Cell Biology | 283 | 10% | 0% | 120 |
| 8 | Biotechnology & Applied Microbiology | 109 | 5% | 0% | 68 |
| 9 | Mycology | 50 | 1% | 0% | 12 |
| 10 | Mathematical & Computational Biology | 29 | 1% | 0% | 15 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ARRHENIUS S E5 | 63158 | 0% | 43% | 6 |
| 2 | PHYSIOCOCHEM MED | 49128 | 0% | 100% | 2 |
| 3 | ALLEGEMEINE MIKROBIOL | 24564 | 0% | 100% | 1 |
| 4 | CANTO BLANCO | 24564 | 0% | 100% | 1 |
| 5 | CLIN SCI GENE GENOME EVOLUT GRP | 24564 | 0% | 100% | 1 |
| 6 | CNRSUMR 6540 DIMARSTN MARINE ENDOUME | 24564 | 0% | 100% | 1 |
| 7 | COMPARAT SEQUENCE ANAL GRP | 24564 | 0% | 100% | 1 |
| 8 | DEP BIOPATOL UMANA | 24564 | 0% | 100% | 1 |
| 9 | ECOL EVOLUT BIOL U43 | 24564 | 0% | 100% | 1 |
| 10 | GENET AL UJAZDOWSKIE 4 | 24564 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEIC ACIDS RESEARCH | 38973 | 20% | 1% | 250 |
| 2 | ZENTRALBLATT FUR BAKTERIOLOGIE MIKROBIOLOGIE UND HYGIENE I ABTEILUNG ORIGINALE C-ALLGEMEINE ANGEWANDTE UND OKOLOGISCHE MIKROBIOLOGIE | 31910 | 1% | 10% | 13 |
| 3 | CELL NUCLEUS | 28517 | 0% | 19% | 6 |
| 4 | SYSTEMATIC AND APPLIED MICROBIOLOGY | 19198 | 4% | 2% | 44 |
| 5 | JOURNAL OF MOLECULAR EVOLUTION | 18744 | 4% | 1% | 54 |
| 6 | ENDOCYTOBIOSIS AND CELL RESEARCH | 7816 | 1% | 4% | 9 |
| 7 | STUDIA BIOPHYSICA | 3289 | 1% | 1% | 16 |
| 8 | CURRENT GENETICS | 2639 | 2% | 1% | 20 |
| 9 | MOLECULAR BIOLOGY | 2275 | 2% | 0% | 21 |
| 10 | FEBS LETTERS | 2243 | 5% | 0% | 62 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | 5S RRNA | 114959 | 2% | 20% | 23 | Search 5S+RRNA | Search 5S+RRNA |
| 2 | 5S RIBOSOMAL RNA | 87707 | 1% | 24% | 15 | Search 5S+RIBOSOMAL+RNA | Search 5S+RIBOSOMAL+RNA |
| 3 | EXPANSION SEGMENT | 86498 | 1% | 39% | 9 | Search EXPANSION+SEGMENT | Search EXPANSION+SEGMENT |
| 4 | RIBOSOMAL PROTEIN L5 | 73687 | 0% | 50% | 6 | Search RIBOSOMAL+PROTEIN+L5 | Search RIBOSOMAL+PROTEIN+L5 |
| 5 | HIDDEN BREAK | 65502 | 0% | 67% | 4 | Search HIDDEN+BREAK | Search HIDDEN+BREAK |
| 6 | RIBOSOME RECONSTITUTION | 49128 | 0% | 100% | 2 | Search RIBOSOME+RECONSTITUTION | Search RIBOSOME+RECONSTITUTION |
| 7 | COMPENSATORY SLIPPAGE | 32751 | 0% | 67% | 2 | Search COMPENSATORY+SLIPPAGE | Search COMPENSATORY+SLIPPAGE |
| 8 | RIBOSOMAL RNA 18S | 32751 | 0% | 67% | 2 | Search RIBOSOMAL+RNA+18S | Search RIBOSOMAL+RNA+18S |
| 9 | 17 25S RIBOSOMAL RNA INTERGENIC REGION | 24564 | 0% | 100% | 1 | Search 17+25S+RIBOSOMAL+RNA+INTERGENIC+REGION | Search 17+25S+RIBOSOMAL+RNA+INTERGENIC+REGION |
| 10 | 18S RIBOSOMAL RNA EVOLUTION | 24564 | 0% | 100% | 1 | Search 18S+RIBOSOMAL+RNA+EVOLUTION | Search 18S+RIBOSOMAL+RNA+EVOLUTION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ERDMANN, VA , HUYSMANS, E , VANDENBERGHE, A , DEWACHTER, R , (1983) COLLECTION OF PUBLISHED 5S AND 5.8S RIBOSOMAL-RNA SEQUENCES.NUCLEIC ACIDS RESEARCH. VOL. 11. ISSUE 1. P. R105-R133 | 55 | 90% | 45 |
| 2 | BARCISZEWSKA, MZ , ERDMANN, VA , BARCISZEWSKI, J , (1996) RIBOSOMAL S-5 RNA: TERTIARY STRUCTURE AND INTERACTIONS WITH PROTEINS.BIOLOGICAL REVIEWS. VOL. 71. ISSUE 1. P. 1 -25 | 52 | 49% | 9 |
| 3 | SCHNARE, MN , DAMBERGER, SH , GRAY, MW , GUTELL, RR , (1996) COMPREHENSIVE COMPARISON OF STRUCTURAL CHARACTERISTICS IN EUKARYOTIC CYTOPLASMIC LARGE SUBUNIT (23 S-LIKE) RIBOSOMAL RNA.JOURNAL OF MOLECULAR BIOLOGY. VOL. 256. ISSUE 4. P. 701-719 | 42 | 42% | 133 |
| 4 | DELIHAS, N , ANDERSEN, J , SINGHAL, RP , (1984) STRUCTURE, FUNCTION AND EVOLUTION OF 5-S RIBOSOMAL-RNAS.PROGRESS IN NUCLEIC ACID RESEARCH AND MOLECULAR BIOLOGY. VOL. 31. ISSUE . P. 161 -190 | 47 | 78% | 47 |
| 5 | BRUNEL, C , ROMBY, P , WESTHOF, E , EHRESMANN, C , EHRESMANN, B , (1991) 3-DIMENSIONAL MODEL OF ESCHERICHIA-COLI RIBOSOMAL 5-S RNA AS DEDUCED FROM STRUCTURE PROBING IN SOLUTION AND COMPUTER MODELING.JOURNAL OF MOLECULAR BIOLOGY. VOL. 221. ISSUE 1. P. 293-308 | 32 | 71% | 62 |
| 6 | ZHAO, YE , WANG, ZH , XU, Y , WU, LP , HU, L , (2013) SECONDARY STRUCTURE PREDICTION FOR COMPLETE RDNA SEQUENCES (18S, 5.8S, AND 28S RDNA) OF DEMODEX FOLLICULORUM, AND COMPARISON OF DIVERGENT DOMAINS STRUCTURES ACROSS ACARI.EXPERIMENTAL PARASITOLOGY. VOL. 135. ISSUE 2. P. 370-381 | 24 | 47% | 3 |
| 7 | SINGHAL, RP , SHAW, JK , (1983) PROKARYOTIC AND EUKARYOTIC 5-S RNAS - PRIMARY SEQUENCES AND PROPOSED SECONDARY STRUCTURES.PROGRESS IN NUCLEIC ACID RESEARCH AND MOLECULAR BIOLOGY. VOL. 28. ISSUE . P. 177 -+ | 37 | 93% | 9 |
| 8 | DELIHAS, N , ANDERSEN, J , (1982) GENERALIZED STRUCTURES OF THE 5S RIBOSOMAL-RNAS.NUCLEIC ACIDS RESEARCH. VOL. 10. ISSUE 22. P. 7323-7344 | 36 | 84% | 95 |
| 9 | NAZAR, RN , (1984) THE RIBOSOMAL 5.8S RNA - EUKARYOTIC ADAPTATION OR PROCESSING VARIANT.CANADIAN JOURNAL OF BIOCHEMISTRY AND CELL BIOLOGY. VOL. 62. ISSUE 6. P. 311-320 | 29 | 97% | 16 |
| 10 | TRIFONOV, EN , BOLSHOI, G , (1983) OPEN AND CLOSED 5-S RIBOSOMAL-RNA, THE ONLY 2 UNIVERSAL STRUCTURES ENCODED IN THE NUCLEOTIDE-SEQUENCES.JOURNAL OF MOLECULAR BIOLOGY. VOL. 169. ISSUE 1. P. 1-13 | 29 | 100% | 26 |
Classes with closest relation at Level 1 |