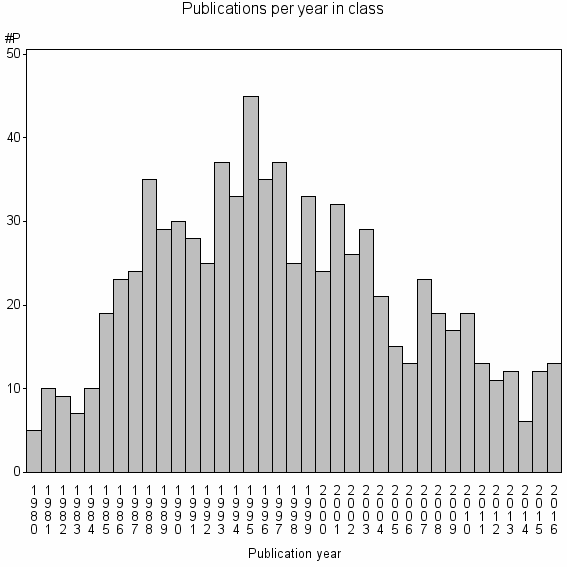
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 13926 | 804 | 39.7 | 75% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 183 | 3 | APTAMER//G QUADRUPLEX//RIBOZYME | 56307 |
| 389 | 2 | NUCLEOSIDES NUCLEOTIDES & NUCLEIC ACIDS//OLIGONUCLEOTIDES//PEPTIDE NUCLEIC ACID | 16949 |
| 13926 | 1 | BIOL MACROMOL STRUCT//SHEARED G CENTER DOT A PAIR//SHEARED A CENTER DOT C PAIR | 804 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | BIOL MACROMOL STRUCT | address | 218740 | 1% | 48% | 12 |
| 2 | SHEARED G CENTER DOT A PAIR | authKW | 189889 | 1% | 100% | 5 |
| 3 | SHEARED A CENTER DOT C PAIR | authKW | 113934 | 0% | 100% | 3 |
| 4 | SHEARED GA PAIRING | authKW | 113934 | 0% | 100% | 3 |
| 5 | G CENTER DOT A MISMATCH | authKW | 85449 | 0% | 75% | 3 |
| 6 | 2 5 LINKED RNA | authKW | 75956 | 0% | 100% | 2 |
| 7 | A FORM CONFORMATION | authKW | 75956 | 0% | 100% | 2 |
| 8 | BASE INTERCALATED DUPLEX | authKW | 75956 | 0% | 100% | 2 |
| 9 | NUCLEIC ACID ENERGETICS | authKW | 75956 | 0% | 100% | 2 |
| 10 | SINGLE RESIDUE LOOP | authKW | 75956 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 6815 | 74% | 0% | 594 |
| 2 | Biophysics | 2342 | 20% | 0% | 162 |
| 3 | Chemistry, Multidisciplinary | 149 | 12% | 0% | 94 |
| 4 | Biochemical Research Methods | 118 | 5% | 0% | 40 |
| 5 | Crystallography | 53 | 3% | 0% | 23 |
| 6 | Chemistry, Physical | 31 | 8% | 0% | 61 |
| 7 | Spectroscopy | 26 | 2% | 0% | 18 |
| 8 | Physics, Atomic, Molecular & Chemical | 20 | 3% | 0% | 27 |
| 9 | Chemistry, Organic | 5 | 3% | 0% | 24 |
| 10 | Cell Biology | 2 | 3% | 0% | 25 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOL MACROMOL STRUCT | 218740 | 1% | 48% | 12 |
| 2 | ARBEITSGRP MAKROMOL STRUKTURANAL | 48825 | 0% | 43% | 3 |
| 3 | ARTHUR AMOS NOYES CHEM | 37978 | 0% | 100% | 1 |
| 4 | BIOCHEM MONTREAL JOINT STRUCT BIOL | 37978 | 0% | 100% | 1 |
| 5 | BIOL STRUCT PHARMACOL MOL | 37978 | 0% | 100% | 1 |
| 6 | BIRCK NANOTCCHNOL | 37978 | 0% | 100% | 1 |
| 7 | BSC CRG IRB JOINT PROGRAM COMPUTAT BIOL | 37978 | 0% | 100% | 1 |
| 8 | CBSE 235 | 37978 | 0% | 100% | 1 |
| 9 | CSSB CHIM STRUCT BIOMOL | 37978 | 0% | 100% | 1 |
| 10 | DESYARBEITSGRP MAKROMOL STRUKTURANAL | 37978 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 15669 | 4% | 1% | 36 |
| 2 | NUCLEIC ACIDS RESEARCH | 12462 | 14% | 0% | 114 |
| 3 | BIOCHEMISTRY | 11653 | 16% | 0% | 128 |
| 4 | BIOPOLYMERS | 7541 | 4% | 1% | 36 |
| 5 | JOURNAL OF MOLECULAR BIOLOGY | 4876 | 7% | 0% | 55 |
| 6 | NUCLEOSIDES NUCLEOTIDES & NUCLEIC ACIDS | 3562 | 3% | 0% | 21 |
| 7 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY | 2469 | 2% | 0% | 17 |
| 8 | NUCLEOSIDES & NUCLEOTIDES | 1418 | 1% | 1% | 5 |
| 9 | JOURNAL OF BIOMOLECULAR NMR | 1140 | 1% | 0% | 9 |
| 10 | RNA | 944 | 1% | 0% | 8 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SHEARED G CENTER DOT A PAIR | 189889 | 1% | 100% | 5 | Search SHEARED+G+CENTER+DOT+A+PAIR | Search SHEARED+G+CENTER+DOT+A+PAIR |
| 2 | SHEARED A CENTER DOT C PAIR | 113934 | 0% | 100% | 3 | Search SHEARED+A+CENTER+DOT+C+PAIR | Search SHEARED+A+CENTER+DOT+C+PAIR |
| 3 | SHEARED GA PAIRING | 113934 | 0% | 100% | 3 | Search SHEARED+GA+PAIRING | Search SHEARED+GA+PAIRING |
| 4 | G CENTER DOT A MISMATCH | 85449 | 0% | 75% | 3 | Search G+CENTER+DOT+A+MISMATCH | Search G+CENTER+DOT+A+MISMATCH |
| 5 | 2 5 LINKED RNA | 75956 | 0% | 100% | 2 | Search 2+5+LINKED+RNA | Search 2+5+LINKED+RNA |
| 6 | A FORM CONFORMATION | 75956 | 0% | 100% | 2 | Search A+FORM+CONFORMATION | Search A+FORM+CONFORMATION |
| 7 | BASE INTERCALATED DUPLEX | 75956 | 0% | 100% | 2 | Search BASE+INTERCALATED+DUPLEX | Search BASE+INTERCALATED+DUPLEX |
| 8 | NUCLEIC ACID ENERGETICS | 75956 | 0% | 100% | 2 | Search NUCLEIC+ACID+ENERGETICS | Search NUCLEIC+ACID+ENERGETICS |
| 9 | SINGLE RESIDUE LOOP | 75956 | 0% | 100% | 2 | Search SINGLE+RESIDUE+LOOP | Search SINGLE+RESIDUE+LOOP |
| 10 | UNPAIRED G STACKS | 75956 | 0% | 100% | 2 | Search UNPAIRED+G+STACKS | Search UNPAIRED+G+STACKS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | NAYAK, RK , VAN ORDEN, A , (2013) COUNTERION AND POLYTHYMIDINE LOOP-LENGTH-DEPENDENT FOLDING AND THERMODYNAMIC STABILITY OF DNA HAIRPINS REVEAL THE UNUSUAL COUNTERION-DEPENDENT STABILITY OF TETRALOOP HAIRPINS.JOURNAL OF PHYSICAL CHEMISTRY B. VOL. 117. ISSUE 45. P. 13956-13966 | 47 | 62% | 2 |
| 2 | CHAKRABORTY, D , COLLEPARDO-GUEVARA, R , WALES, DJ , (2014) ENERGY LANDSCAPES, FOLDING MECHANISMS, AND KINETICS OF RNA TETRA LOOP HAIRPINS.JOURNAL OF THE AMERICAN CHEMICAL SOCIETY. VOL. 136. ISSUE 52. P. 18052 -18061 | 37 | 39% | 10 |
| 3 | LAM, SL , CHI, LM , (2010) USE OF CHEMICAL SHIFTS FOR STRUCTURAL STUDIES OF NUCLEIC ACIDS.PROGRESS IN NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY. VOL. 56. ISSUE 3. P. 289-310 | 47 | 34% | 21 |
| 4 | CHOU, SH , CHIN, KH , WANG, AHJ , (2003) UNUSUAL DNA DUPLEX AND HAIRPIN MOTIFS.NUCLEIC ACIDS RESEARCH. VOL. 31. ISSUE 10. P. 2461-2474 | 47 | 40% | 59 |
| 5 | MINER, JC , CHEN, AA , GARCIA, AE , (2016) FREE-ENERGY LANDSCAPE OF A HYPERSTABLE RNA TETRALOOP.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 113. ISSUE 24. P. 6665 -6670 | 27 | 47% | 2 |
| 6 | CHANG, CY , STELLWAGEN, NC , (2011) TANDEM GA RESIDUES ON OPPOSITE SIDES OF THE LOOP IN MOLECULAR BEACON-LIKE DNA HAIRPINS COMPACT THE LOOP AND INCREASE HAIRPIN STABILITY.BIOCHEMISTRY. VOL. 50. ISSUE 42. P. 9148-9157 | 31 | 48% | 3 |
| 7 | VANDONGEN, MJP , MOOREN, MMW , WILLEMS, EFA , VANDERMAREL, GA , VANBOOM, JH , WIJMENGA, SS , HILBERS, CW , (1997) STRUCTURAL FEATURES OF THE DNA HAIRPIN D(ATCCTA-GTTA-TAGGAT): FORMATION OF A G-A BASE PAIR IN THE LOOP.NUCLEIC ACIDS RESEARCH. VOL. 25. ISSUE 8. P. 1537-1547 | 27 | 71% | 43 |
| 8 | KUHROVA, P , BANAS, P , BEST, RB , SPONER, J , OTYEPKA, M , (2013) COMPUTER FOLDING OF RNA TETRALOOPS? ARE WE THERE YET?.JOURNAL OF CHEMICAL THEORY AND COMPUTATION. VOL. 9. ISSUE 4. P. 2115-2125 | 32 | 35% | 22 |
| 9 | MELNYKOV, AV , NAYAK, RK , HALL, KB , VAN ORDEN, A , (2015) EFFECT OF LOOP COMPOSITION ON THE STABILITY AND FOLDING KINETICS OF RNA HAIRPINS WITH LARGE LOOPS.BIOCHEMISTRY. VOL. 54. ISSUE 10. P. 1886 -1896 | 21 | 57% | 1 |
| 10 | CHEN, AA , GARCIA, AE , (2013) HIGH-RESOLUTION REVERSIBLE FOLDING OF HYPERSTABLE RNA TETRALOOPS USING MOLECULAR DYNAMICS SIMULATIONS.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 110. ISSUE 42. P. 16820-16825 | 18 | 47% | 72 |
Classes with closest relation at Level 1 |