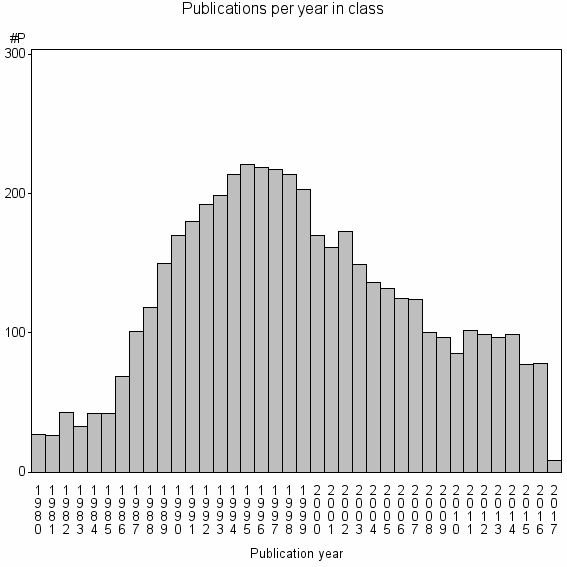
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2254 | 4692 | 42.3 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 328 | 3 | PHOSPHOLIPASE D//STIM1//REGUCALCIN | 38118 |
| 2254 | 2 | PHOSPHOLIPASE D//DIACYLGLYCEROL KINASE//PHOSPHATIDIC ACID | 4692 |
| 3408 | 1 | PHOSPHOLIPASE D//PHOSPHATIDIC ACID//PHOSPHOLIPASE D1 | 1990 |
| 6896 | 1 | PHOSPHOLIPASE C DELTA 1//GENOME BIOSIGNAL//PHOSPHOLIPASE C | 1431 |
| 16564 | 1 | CELLULAR SIGNALLING//DNA BOUND LIPIDS//PI PLCBETA1 | 647 |
| 20059 | 1 | DIACYLGLYCEROL KINASE//DIACYLGLYCEROL KINASE DGK//PHOSPHATIDIC ACID | 478 |
| 31931 | 1 | BAYERS JUNCTIONS//DNA MEMBRANE COMPLEXES//DNA MEMBRANE CONTACTS | 146 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PHOSPHOLIPASE D | authKW | 1812138 | 13% | 46% | 606 |
| 2 | DIACYLGLYCEROL KINASE | authKW | 624068 | 3% | 63% | 153 |
| 3 | PHOSPHATIDIC ACID | authKW | 575361 | 5% | 35% | 251 |
| 4 | PHOSPHOLIPASE C | authKW | 192712 | 5% | 12% | 252 |
| 5 | PHOSPHOLIPASE D1 | authKW | 115663 | 1% | 66% | 27 |
| 6 | PHOSPHOLIPASE D2 | authKW | 108573 | 0% | 76% | 22 |
| 7 | PHOSPHOLIPASE C DELTA 1 | authKW | 86977 | 0% | 70% | 19 |
| 8 | DIACYLGLYCEROL | authKW | 86962 | 3% | 10% | 130 |
| 9 | PHOSPHATIDYLETHANOL | authKW | 78755 | 1% | 34% | 36 |
| 10 | PLD1 | authKW | 78061 | 0% | 67% | 18 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 26966 | 62% | 0% | 2900 |
| 2 | Cell Biology | 12554 | 27% | 0% | 1286 |
| 3 | Biophysics | 9125 | 17% | 0% | 783 |
| 4 | CYTOLOGY & HISTOLOGY | 1517 | 1% | 1% | 43 |
| 5 | Physiology | 167 | 3% | 0% | 146 |
| 6 | Oncology | 126 | 5% | 0% | 233 |
| 7 | Pharmacology & Pharmacy | 107 | 6% | 0% | 264 |
| 8 | Neurosciences | 88 | 5% | 0% | 251 |
| 9 | Medicine, Research & Experimental | 82 | 3% | 0% | 132 |
| 10 | Hematology | 35 | 2% | 0% | 80 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | GENOME BIOSIGNAL | 76696 | 1% | 47% | 25 |
| 2 | STN ABORAT | 48908 | 0% | 40% | 19 |
| 3 | HUMAN ANAT SCI | 41948 | 0% | 32% | 20 |
| 4 | CELLULAR SIGNALLING | 37678 | 1% | 16% | 37 |
| 5 | DIPARTIMENTO SCI ANAT UMANE FISIOPATOL PARATO | 31287 | 0% | 25% | 19 |
| 6 | DIPARTIMENTO MORFOL UMANA NORMALE | 26004 | 0% | 29% | 14 |
| 7 | NUCL LIPID BIOPATHOL | 24516 | 0% | 54% | 7 |
| 8 | HUMAN GENETSEONGBUK KU | 20818 | 0% | 80% | 4 |
| 9 | CITOMORFOL NORMALE PATOL | 20328 | 0% | 63% | 5 |
| 10 | CLIN EXPT MED PHYSIOPATHOL | 19518 | 0% | 100% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ADVANCES IN ENZYME REGULATION | 20104 | 1% | 7% | 43 |
| 2 | CELLULAR SIGNALLING | 17599 | 2% | 3% | 109 |
| 3 | BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR AND CELL BIOLOGY OF LIPIDS | 15789 | 2% | 3% | 81 |
| 4 | JOURNAL OF BIOLOGICAL CHEMISTRY | 15220 | 13% | 0% | 606 |
| 5 | BIOCHEMICAL JOURNAL | 11817 | 5% | 1% | 233 |
| 6 | JOURNAL OF LIPID MEDIATORS AND CELL SIGNALLING | 6756 | 0% | 6% | 17 |
| 7 | EXPERIMENTAL AND MOLECULAR MEDICINE | 5279 | 1% | 2% | 34 |
| 8 | BIOCHEMICAL AND BIOPHYSICAL RESEARCH COMMUNICATIONS | 5000 | 5% | 0% | 235 |
| 9 | FEBS LETTERS | 4486 | 4% | 0% | 172 |
| 10 | BIOCHIMICA ET BIOPHYSICA ACTA-LIPIDS AND LIPID METABOLISM | 2892 | 0% | 2% | 22 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PHOSPHOLIPASE D | 1812138 | 13% | 46% | 606 | Search PHOSPHOLIPASE+D | Search PHOSPHOLIPASE+D |
| 2 | DIACYLGLYCEROL KINASE | 624068 | 3% | 63% | 153 | Search DIACYLGLYCEROL+KINASE | Search DIACYLGLYCEROL+KINASE |
| 3 | PHOSPHATIDIC ACID | 575361 | 5% | 35% | 251 | Search PHOSPHATIDIC+ACID | Search PHOSPHATIDIC+ACID |
| 4 | PHOSPHOLIPASE C | 192712 | 5% | 12% | 252 | Search PHOSPHOLIPASE+C | Search PHOSPHOLIPASE+C |
| 5 | PHOSPHOLIPASE D1 | 115663 | 1% | 66% | 27 | Search PHOSPHOLIPASE+D1 | Search PHOSPHOLIPASE+D1 |
| 6 | PHOSPHOLIPASE D2 | 108573 | 0% | 76% | 22 | Search PHOSPHOLIPASE+D2 | Search PHOSPHOLIPASE+D2 |
| 7 | PHOSPHOLIPASE C DELTA 1 | 86977 | 0% | 70% | 19 | Search PHOSPHOLIPASE+C+DELTA+1 | Search PHOSPHOLIPASE+C+DELTA+1 |
| 8 | DIACYLGLYCEROL | 86962 | 3% | 10% | 130 | Search DIACYLGLYCEROL | Search DIACYLGLYCEROL |
| 9 | PHOSPHATIDYLETHANOL | 78755 | 1% | 34% | 36 | Search PHOSPHATIDYLETHANOL | Search PHOSPHATIDYLETHANOL |
| 10 | PLD1 | 78061 | 0% | 67% | 18 | Search PLD1 | Search PLD1 |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | MCDERMOTT, M , WAKELAM, MJO , MORRIS, AJ , (2004) PHOSPHOLIPASE D.BIOCHEMISTRY AND CELL BIOLOGY. VOL. 82. ISSUE 1. P. 225 -253 | 216 | 68% | 233 |
| 2 | EXTON, JH , (2002) PHOSPHOLIPASE D - STRUCTURE, REGULATION AND FUNCTION.REVIEWS OF PHYSIOLOGY BIOCHEMISTRY AND PHARMACOLOGY, VOL 144. VOL. 144. ISSUE . P. 1 -94 | 236 | 64% | 160 |
| 3 | BRUNTZ, RC , LINDSLEY, CW , BROWN, HA , (2014) PHOSPHOLIPASE D SIGNALING PATHWAYS AND PHOSPHATIDIC ACID AS THERAPEUTIC TARGETS IN CANCER.PHARMACOLOGICAL REVIEWS. VOL. 66. ISSUE 4. P. 1033 -1079 | 254 | 42% | 16 |
| 4 | GOMEZ-CAMBRONERO, J , (2011) THE EXQUISITE REGULATION OF PLD2 BY A WEALTH OF INTERACTING PROTEINS: S6K, GRB2, SOS, WASP AND RAC2 (AND A SURPRISE DISCOVERY: PLD2 IS A GEF).CELLULAR SIGNALLING. VOL. 23. ISSUE 12. P. 1885 -1895 | 157 | 79% | 7 |
| 5 | REBECCHI, MJ , PENTYALA, SN , (2000) STRUCTURE, FUNCTION, AND CONTROL OF PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C.PHYSIOLOGICAL REVIEWS. VOL. 80. ISSUE 4. P. 1291 -1335 | 191 | 46% | 679 |
| 6 | JENKINS, GM , FROHMAN, MA , (2005) PHOSPHOLIPASE D: A LIPID CENTRIC REVIEW.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 62. ISSUE 19-20. P. 2305-2316 | 122 | 78% | 267 |
| 7 | SINGER, WD , BROWN, HA , STERNWEIS, PC , (1997) REGULATION OF EUKARYOTIC PHOSPHATIDYLINOSITOL-SPECIFIC PHOSPHOLIPASE C AND PHOSPHOLIPASE D.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 66. ISSUE . P. 475 -509 | 188 | 61% | 289 |
| 8 | LISCOVITCH, M , CZARNY, M , FIUCCI, G , TANG, XQ , (2000) PHOSPHOLIPASE D: MOLECULAR AND CELL BIOLOGY OF A NOVEL GENE FAMILY.BIOCHEMICAL JOURNAL. VOL. 345. ISSUE . P. 401 -415 | 145 | 66% | 391 |
| 9 | KADAMUR, G , ROSS, EM , (2013) MAMMALIAN PHOSPHOLIPASE C.ANNUAL REVIEW OF PHYSIOLOGY, VOL 75. VOL. 75. ISSUE . P. 127-154 | 113 | 64% | 69 |
| 10 | SHULGA, YV , TOPHAM, MK , EPAND, RM , (2011) REGULATION AND FUNCTIONS OF DIACYLGLYCEROL KINASES.CHEMICAL REVIEWS. VOL. 111. ISSUE 10. P. 6186 -6208 | 141 | 57% | 56 |
Classes with closest relation at Level 2 |