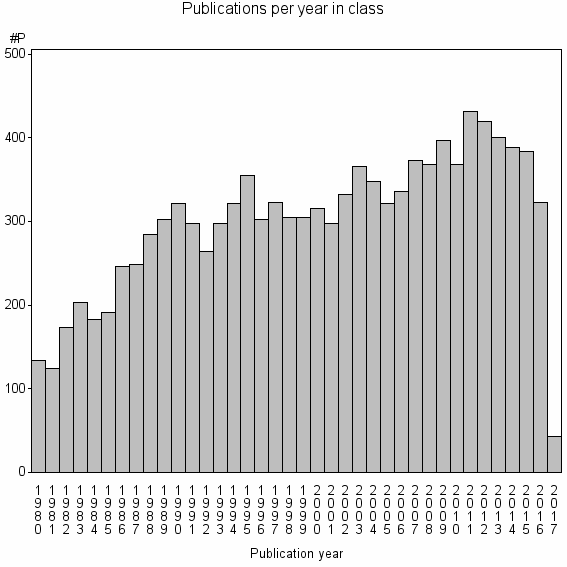
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 892 | 11392 | 45.3 | 82% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 7 | 4 | INFECTIOUS DISEASES//MICROBIOLOGY//VIROLOGY | 1353914 |
| 33 | 3 | JOURNAL OF BACTERIOLOGY//MICROBIOLOGY//MOLECULAR MICROBIOLOGY | 107033 |
| 892 | 2 | PROTEIN TRANSLOCATION//PROTEIN IMPORT//SECA | 11392 |
| 2349 | 1 | SECA//SECB//SECYEG | 2274 |
| 2667 | 1 | PROTEIN IMPORT//TOM COMPLEX//MITOCHONDRIAL PROTEIN IMPORT | 2179 |
| 4955 | 1 | SIGNAL RECOGNITION PARTICLE//TRANSLOCON//FFH | 1697 |
| 5827 | 1 | PORIN//MALTOPORIN//OUTER MEMBRANE PROTEIN | 1570 |
| 9219 | 1 | CHLOROPLAST PROTEIN IMPORT//PROTEIN IMPORT//CHLOROPLAST | 1176 |
| 12801 | 1 | AUTOTRANSPORTER//AUTODISPLAY//BAM COMPLEX | 880 |
| 15103 | 1 | HTRA1//HTRA//HTRA2 | 731 |
| 15877 | 1 | TWIN ARGININE TRANSLOCATION//TAT PATHWAY//TWIN ARGININE | 686 |
| 28802 | 1 | LOLB//LOLA//LOL SYSTEM | 199 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PROTEIN TRANSLOCATION | authKW | 528472 | 3% | 60% | 328 |
| 2 | PROTEIN IMPORT | authKW | 451311 | 3% | 59% | 288 |
| 3 | SECA | authKW | 292417 | 1% | 82% | 133 |
| 4 | SIGNAL RECOGNITION PARTICLE | authKW | 291593 | 1% | 78% | 140 |
| 5 | PORIN | authKW | 205687 | 2% | 36% | 211 |
| 6 | PROTEIN TRANSPORT | authKW | 174323 | 2% | 38% | 172 |
| 7 | AUTOTRANSPORTER | authKW | 169678 | 1% | 71% | 89 |
| 8 | TRANSLOCON | authKW | 147665 | 1% | 65% | 85 |
| 9 | PROTEIN TARGETING | authKW | 123793 | 1% | 29% | 159 |
| 10 | TWIN ARGININE TRANSLOCATION | authKW | 122212 | 0% | 89% | 51 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 68382 | 63% | 0% | 7182 |
| 2 | Cell Biology | 26056 | 26% | 0% | 2907 |
| 3 | Microbiology | 17975 | 18% | 0% | 2005 |
| 4 | Biophysics | 11667 | 12% | 0% | 1418 |
| 5 | Plant Sciences | 1050 | 6% | 0% | 664 |
| 6 | Biochemical Research Methods | 587 | 3% | 0% | 385 |
| 7 | Biotechnology & Applied Microbiology | 288 | 4% | 0% | 422 |
| 8 | Genetics & Heredity | 181 | 3% | 0% | 364 |
| 9 | Biology | 140 | 2% | 0% | 198 |
| 10 | Developmental Biology | 12 | 1% | 0% | 68 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ZBMZ | 113086 | 1% | 50% | 84 |
| 2 | ZENTRUM BIOCHEM MOL ZELLFOR | 38809 | 0% | 50% | 29 |
| 3 | FAK BIOL | 24836 | 1% | 9% | 108 |
| 4 | CANADA CHAIR BACTERIAL ANIM DIS | 24348 | 0% | 91% | 10 |
| 5 | GENET BIOCHEM BRANCH | 21960 | 0% | 21% | 39 |
| 6 | MOL MICROBIOL | 21414 | 2% | 4% | 225 |
| 7 | GRONINGEN BIOMOL SCI BIOTECHNOL | 18196 | 1% | 6% | 113 |
| 8 | MEMBRANE PROTE | 15719 | 0% | 28% | 21 |
| 9 | FB BIOL GEOG | 15490 | 0% | 64% | 9 |
| 10 | BIOCHEM HERMANN HERDER STR 7 | 14280 | 0% | 67% | 8 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BACTERIOLOGY | 39440 | 6% | 2% | 706 |
| 2 | JOURNAL OF BIOLOGICAL CHEMISTRY | 33030 | 12% | 1% | 1394 |
| 3 | EMBO JOURNAL | 21627 | 3% | 2% | 376 |
| 4 | MOLECULAR MICROBIOLOGY | 14609 | 2% | 2% | 258 |
| 5 | FEBS LETTERS | 10643 | 4% | 1% | 413 |
| 6 | JOURNAL OF CELL BIOLOGY | 10371 | 2% | 2% | 253 |
| 7 | JOURNAL OF MOLECULAR BIOLOGY | 9809 | 3% | 1% | 299 |
| 8 | CELL | 5824 | 2% | 1% | 185 |
| 9 | BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH | 5517 | 1% | 2% | 98 |
| 10 | EUROPEAN JOURNAL OF BIOCHEMISTRY | 4831 | 2% | 1% | 195 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PROTEIN TRANSLOCATION | 528472 | 3% | 60% | 328 | Search PROTEIN+TRANSLOCATION | Search PROTEIN+TRANSLOCATION |
| 2 | PROTEIN IMPORT | 451311 | 3% | 59% | 288 | Search PROTEIN+IMPORT | Search PROTEIN+IMPORT |
| 3 | SECA | 292417 | 1% | 82% | 133 | Search SECA | Search SECA |
| 4 | SIGNAL RECOGNITION PARTICLE | 291593 | 1% | 78% | 140 | Search SIGNAL+RECOGNITION+PARTICLE | Search SIGNAL+RECOGNITION+PARTICLE |
| 5 | PORIN | 205687 | 2% | 36% | 211 | Search PORIN | Search PORIN |
| 6 | PROTEIN TRANSPORT | 174323 | 2% | 38% | 172 | Search PROTEIN+TRANSPORT | Search PROTEIN+TRANSPORT |
| 7 | AUTOTRANSPORTER | 169678 | 1% | 71% | 89 | Search AUTOTRANSPORTER | Search AUTOTRANSPORTER |
| 8 | TRANSLOCON | 147665 | 1% | 65% | 85 | Search TRANSLOCON | Search TRANSLOCON |
| 9 | PROTEIN TARGETING | 123793 | 1% | 29% | 159 | Search PROTEIN+TARGETING | Search PROTEIN+TARGETING |
| 10 | TWIN ARGININE TRANSLOCATION | 122212 | 0% | 89% | 51 | Search TWIN+ARGININE+TRANSLOCATION | Search TWIN+ARGININE+TRANSLOCATION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | KUDVA, R , DENKS, K , KUHN, P , VOGT, A , MULLER, M , KOCH, HG , (2013) PROTEIN TRANSLOCATION ACROSS THE INNER MEMBRANE OF GRAM-NEGATIVE BACTERIA: THE SEC AND TAT DEPENDENT PROTEIN TRANSPORT PATHWAYS.RESEARCH IN MICROBIOLOGY. VOL. 164. ISSUE 6. P. 505 -534 | 357 | 94% | 39 |
| 2 | DENKS, K , VOGT, A , SACHELARU, I , PETRIMAN, NA , KUDVA, R , KOCH, HG , (2014) THE SEC TRANSLOCON MEDIATED PROTEIN TRANSPORT IN PROKARYOTES AND EUKARYOTES.MOLECULAR MEMBRANE BIOLOGY. VOL. 31. ISSUE 2-3. P. 58 -84 | 342 | 84% | 28 |
| 3 | AKOPIAN, D , SHEN, K , ZHANG, X , SHAN, SO , (2013) SIGNAL RECOGNITION PARTICLE: AN ESSENTIAL PROTEIN-TARGETING MACHINE.ANNUAL REVIEW OF BIOCHEMISTRY, VOL 82. VOL. 82. ISSUE . P. 693-721 | 205 | 94% | 90 |
| 4 | NEUPERT, W , (1997) PROTEIN IMPORT INTO MITOCHONDRIA.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 66. ISSUE . P. 863-917 | 320 | 68% | 831 |
| 5 | DU PLESSIS, DJF , NOUWEN, N , DRIESSEN, AJM , (2011) THE SEC TRANSLOCASE.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1808. ISSUE 3. P. 851-865 | 229 | 90% | 115 |
| 6 | NATALE, P , BRUSER, T , DRIESSEN, AJM , (2008) SEC- AND TAT-MEDIATED PROTEIN SECRETION ACROSS THE BACTERIAL CYTOPLASMIC MEMBRANE - DISTINCT TRANSLOCASES AND MECHANISMS.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1778. ISSUE 9. P. 1735-1756 | 245 | 90% | 156 |
| 7 | PUGSLEY, AP , (1993) THE COMPLETE GENERAL SECRETORY PATHWAY IN GRAM-NEGATIVE BACTERIA.MICROBIOLOGICAL REVIEWS. VOL. 57. ISSUE 1. P. 50 -108 | 328 | 58% | 1544 |
| 8 | SHI, LX , THEG, SM , (2013) THE CHLOROPLAST PROTEIN IMPORT SYSTEM: FROM ALGAE TO TREES.BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH. VOL. 1833. ISSUE 2. P. 314-331 | 193 | 89% | 54 |
| 9 | NEUPERT, W , HERRMANN, JM , (2007) TRANSLOCATION OF PROTEINS INTO MITOCHONDRIA.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 76. ISSUE . P. 723 -749 | 154 | 83% | 681 |
| 10 | HOHR, AIC , STRAUB, SP , WARSCHEID, B , BECKER, T , WIEDEMANN, N , (2015) ASSEMBLY OF BETA-BARREL PROTEINS IN THE MITOCHONDRIAL OUTER MEMBRANE.BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH. VOL. 1853. ISSUE 1. P. 74 -88 | 219 | 81% | 12 |
Classes with closest relation at Level 2 |