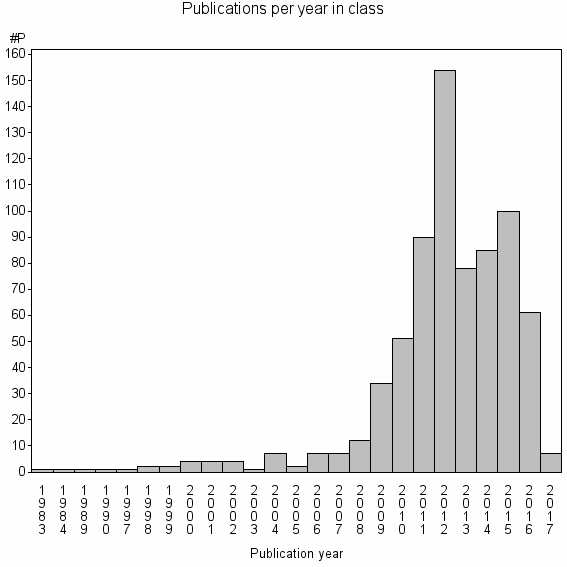
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 15359 | 717 | 31.9 | 78% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 15359 | 1 | STANDARDS IN GENOMIC SCIENCES//BIOL DATA MANAGEMENT TECHNOL//GEBA | 717 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | STANDARDS IN GENOMIC SCIENCES | journal | 6713367 | 45% | 49% | 325 |
| 2 | BIOL DATA MANAGEMENT TECHNOL | address | 5098157 | 24% | 69% | 173 |
| 3 | GEBA | authKW | 2986402 | 13% | 75% | 94 |
| 4 | TAXONO GENOMICS | authKW | 1415171 | 5% | 92% | 36 |
| 5 | NON MOTILE | authKW | 1197707 | 6% | 63% | 45 |
| 6 | CHEMOORGANOTROPHIC | authKW | 1126346 | 5% | 73% | 36 |
| 7 | MOTILE | authKW | 1060226 | 5% | 66% | 38 |
| 8 | CULTUROMICS | authKW | 1042255 | 5% | 64% | 38 |
| 9 | STRICTLY ANAEROBIC | authKW | 627174 | 3% | 82% | 18 |
| 10 | ROOT NODULE BACTERIA | authKW | 543375 | 5% | 36% | 35 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Microbiology | 25162 | 77% | 0% | 552 |
| 2 | Genetics & Heredity | 11571 | 52% | 0% | 376 |
| 3 | Biotechnology & Applied Microbiology | 540 | 13% | 0% | 92 |
| 4 | Mathematical & Computational Biology | 117 | 3% | 0% | 19 |
| 5 | Biochemical Research Methods | 28 | 3% | 0% | 22 |
| 6 | Infectious Diseases | 1 | 1% | 0% | 9 |
| 7 | Biochemistry & Molecular Biology | 1 | 6% | 0% | 45 |
| 8 | Evolutionary Biology | 0 | 1% | 0% | 4 |
| 9 | Virology | 0 | 1% | 0% | 5 |
| 10 | Gastroenterology & Hepatology | 0 | 1% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOL DATA MANAGEMENT TECHNOL | 5098157 | 24% | 69% | 173 |
| 2 | AUSTRALIAN ECOGEN | 455461 | 5% | 28% | 38 |
| 3 | RHIZOBIUM STUDIES | 337138 | 5% | 23% | 34 |
| 4 | JOINT GENOME | 325342 | 9% | 12% | 62 |
| 5 | GENOME BIOL PROGRAM | 313725 | 3% | 39% | 19 |
| 6 | AUSTRALIAN ECOGENOM | 279694 | 2% | 39% | 17 |
| 7 | ALGORITHM BIOL | 265322 | 3% | 35% | 18 |
| 8 | THEODOSIUS DOBZHANSKY GENOME BIONFORMAT | 247771 | 1% | 73% | 8 |
| 9 | GENOME OURCE | 229430 | 3% | 28% | 19 |
| 10 | PLAGUE BRUCELLOSIS PREVENT CONTROL BASE | 170341 | 1% | 67% | 6 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | STANDARDS IN GENOMIC SCIENCES | 6713367 | 45% | 49% | 325 |
| 2 | MARINE GENOMICS | 48196 | 3% | 5% | 23 |
| 3 | JOURNAL OF BACTERIOLOGY | 22080 | 18% | 0% | 131 |
| 4 | GUT PATHOGENS | 7493 | 1% | 3% | 7 |
| 5 | JOURNAL OF BIOTECHNOLOGY | 7422 | 5% | 1% | 34 |
| 6 | OMICS-A JOURNAL OF INTEGRATIVE BIOLOGY | 4903 | 1% | 1% | 9 |
| 7 | INTERNATIONAL JOURNAL OF SYSTEMATIC AND EVOLUTIONARY MICROBIOLOGY | 2189 | 3% | 0% | 21 |
| 8 | SYSTEMATIC AND APPLIED MICROBIOLOGY | 610 | 1% | 0% | 6 |
| 9 | VIROLOGICA SINICA | 551 | 0% | 1% | 1 |
| 10 | NUCLEIC ACIDS RESEARCH | 534 | 3% | 0% | 23 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GEBA | 2986402 | 13% | 75% | 94 | Search GEBA | Search GEBA |
| 2 | TAXONO GENOMICS | 1415171 | 5% | 92% | 36 | Search TAXONO+GENOMICS | Search TAXONO+GENOMICS |
| 3 | NON MOTILE | 1197707 | 6% | 63% | 45 | Search NON+MOTILE | Search NON+MOTILE |
| 4 | CHEMOORGANOTROPHIC | 1126346 | 5% | 73% | 36 | Search CHEMOORGANOTROPHIC | Search CHEMOORGANOTROPHIC |
| 5 | MOTILE | 1060226 | 5% | 66% | 38 | Search MOTILE | Search MOTILE |
| 6 | CULTUROMICS | 1042255 | 5% | 64% | 38 | Search CULTUROMICS | Search CULTUROMICS |
| 7 | STRICTLY ANAEROBIC | 627174 | 3% | 82% | 18 | Search STRICTLY+ANAEROBIC | Search STRICTLY+ANAEROBIC |
| 8 | ROOT NODULE BACTERIA | 543375 | 5% | 36% | 35 | Search ROOT+NODULE+BACTERIA | Search ROOT+NODULE+BACTERIA |
| 9 | ALPHAPROTEOBACTERIA | 510976 | 6% | 29% | 42 | Search ALPHAPROTEOBACTERIA | Search ALPHAPROTEOBACTERIA |
| 10 | NON SPORULATING | 438027 | 2% | 86% | 12 | Search NON+SPORULATING | Search NON+SPORULATING |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ANGELAKIS, E , BIBI, F , RAMASAMY, D , AZHAR, EI , JIMAN-FATANI, AA , ABOUSHOUSHAH, SM , LAGIER, JC , ROBERT, C , CAPUTO, A , YASIR, M , ET AL (2014) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF CLOSTRIDIUM SAUDII SP NOV.STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 1. P. - | 38 | 79% | 0 |
| 2 | LAGIER, JC , BIBI, F , RAMASAMY, D , AZHAR, EI , ROBERT, C , YASIR, M , JIMAN-FATANI, AA , ALSHALI, KZ , FOURNIER, PE , RAOULT, D , (2014) NON CONTIGUOUS-FINISHED GENOME SEQUENCE AND DESCRIPTION OF CLOSTRIDIUM JEDDAHENSE SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 37 | 80% | 0 |
| 3 | PADMANABHAN, R , DUBOURG, G , LAGIER, JC , NGUYEN, TT , COUDERC, C , ROSSI-TAMISIER, M , CAPUTO, A , RAOULT, D , FOURNIER, PE , (2014) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF COLLINSELLA MASSILIENSIS SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 37 | 74% | 0 |
| 4 | RAMASAMY, D , LAGIER, JC , ROSSI-, M , PFLEIDERER, TA , MICHELLE, C , COUDERC, C , RAOULT, D , FOURNIER, PE , (2014) GENOME SEQUENCE AND DESCRIPTION OF BACTEROIDES TIMONENSIS SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 39 | 70% | 0 |
| 5 | PADMANABHAN, R , LAGIER, JC , DANGUI, NPM , MICHELLE, C , COUDERC, C , RAOULT, D , FOURNIER, PE , (2013) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF MEGASPHAERA MASSILIENSIS SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 8. ISSUE 3. P. 525-538 | 33 | 79% | 1 |
| 6 | EDOUARD, S , SANKAR, S , DANGUI, NPM , LAGIER, JC , MICHELLE, C , RAOULT, D , FOURNIER, PE , (2014) GENOME SEQUENCE AND DESCRIPTION OF NESTERENKONIA MASSILIENSIS SP NOV STRAIN NP1(T).STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 36 | 69% | 0 |
| 7 | EDOUARD, S , BIBI, F , DHAMODHARAN, R , LAGIER, JC , AZHAR, EI , ROBERT, C , CAPUTO, A , YASIR, M , JIMAN-FATANI, AA , ALAWI, M , ET AL (2014) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF CORYNEBACTERIUM JEDDAHENSE SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 37 | 67% | 0 |
| 8 | MISHRA, AK , LAGIER, JC , PFLEIDERER, A , NGUYEN, TT , CAPUTO, A , RAOULT, D , FOURNIER, PE , (2013) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF HOLDEMANIA MASSILIENSIS SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 2. P. 395-409 | 30 | 81% | 2 |
| 9 | PFLEIDERER, A , MISHRA, AK , LAGIER, JC , ROBERT, C , CAPUTO, A , RAOULT, D , FOURNIER, PE , (2014) NON-CONTIGUOUS FINISHED GENOME SEQUENCE AND DESCRIPTION OF ALISTIPES IHUMII SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 30 | 77% | 1 |
| 10 | PADMANABHAN, R , DUBOURG, G , LAGIER, JC , COUDERC, C , MICHELLE, C , RAOULT, D , FOURNIER, PE , (2014) GENOME SEQUENCE AND DESCRIPTION OF CORYNEBACTERIUM IHUMII SP NOV..STANDARDS IN GENOMIC SCIENCES. VOL. 9. ISSUE 3. P. - | 31 | 72% | 0 |
Classes with closest relation at Level 1 |