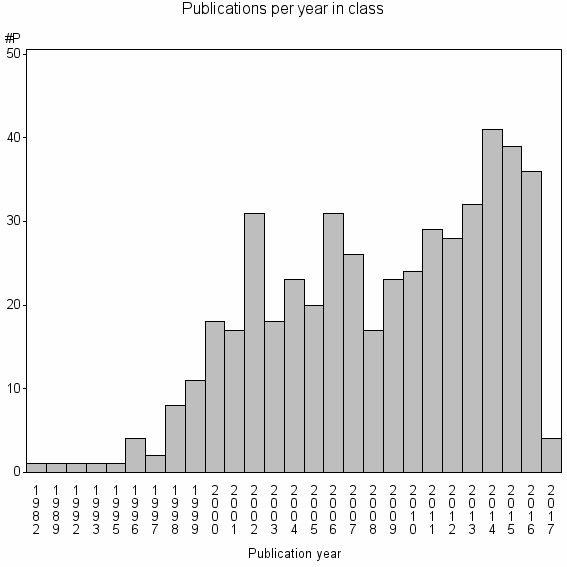
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 19843 | 487 | 44.4 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 389 | 3 | PEROXISOME//CARNITINE//JOURNAL OF INHERITED METABOLIC DISEASE | 30841 |
| 3076 | 2 | PEPTIDE DEFORMYLASE//ENOYL ACP REDUCTASE//ACYL CARRIER PROTEIN | 2395 |
| 19843 | 1 | ESSENTIAL GENES//TN SEQ//SUBTRACTIVE GENOMICS | 487 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ESSENTIAL GENES | authKW | 610103 | 9% | 23% | 43 |
| 2 | TN SEQ | authKW | 236324 | 1% | 54% | 7 |
| 3 | SUBTRACTIVE GENOMICS | authKW | 225715 | 1% | 60% | 6 |
| 4 | MINIMAL GENOME | authKW | 188088 | 2% | 33% | 9 |
| 5 | ALLELIC REPLACEMENT MUTAGENESIS | authKW | 125400 | 0% | 100% | 2 |
| 6 | COMPUTATIONAL COMPARATIVE GENOMICS | authKW | 125400 | 0% | 100% | 2 |
| 7 | ESSENTIAL GENE PREDICTION | authKW | 125400 | 0% | 100% | 2 |
| 8 | ESSENTIAL GENOME | authKW | 125400 | 0% | 100% | 2 |
| 9 | IN SILICO DRUG TARGETS | authKW | 125400 | 0% | 100% | 2 |
| 10 | KAAS SERVER | authKW | 125400 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Microbiology | 3132 | 34% | 0% | 165 |
| 2 | Biotechnology & Applied Microbiology | 1390 | 23% | 0% | 114 |
| 3 | Genetics & Heredity | 784 | 18% | 0% | 86 |
| 4 | Biochemical Research Methods | 467 | 11% | 0% | 54 |
| 5 | Biochemistry & Molecular Biology | 436 | 28% | 0% | 135 |
| 6 | Mathematical & Computational Biology | 341 | 5% | 0% | 25 |
| 7 | Pharmacology & Pharmacy | 67 | 9% | 0% | 46 |
| 8 | Infectious Diseases | 34 | 3% | 0% | 16 |
| 9 | Biophysics | 24 | 4% | 0% | 18 |
| 10 | Biology | 14 | 2% | 0% | 11 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SECTOR AGR MICROBIOL AGRIFOOD TECHNOL | 125400 | 0% | 100% | 2 |
| 2 | SECTOR FOOD TECHNOL | 125400 | 0% | 100% | 2 |
| 3 | STRATEG TRAINING PROGRAM BIOINFORMAT | 125400 | 0% | 100% | 2 |
| 4 | 4MD | 62700 | 0% | 100% | 1 |
| 5 | ANTIBACT | 62700 | 0% | 100% | 1 |
| 6 | ANTIMICROBIALS HOST DEFENCE CEDD | 62700 | 0% | 100% | 1 |
| 7 | BIOINFONNAT SCI TECHNOL | 62700 | 0% | 100% | 1 |
| 8 | CHIM BIOIND | 62700 | 0% | 100% | 1 |
| 9 | FG GARUNGSTECHNOL FERMENTAT TECHNOL | 62700 | 0% | 100% | 1 |
| 10 | GENOME EXP S | 62700 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CURRENT OPINION IN MICROBIOLOGY | 9895 | 3% | 1% | 17 |
| 2 | MBIO | 2276 | 2% | 0% | 8 |
| 3 | TRENDS IN MICROBIOLOGY | 2225 | 2% | 0% | 8 |
| 4 | JOURNAL OF BIOINFORMATICS AND COMPUTATIONAL BIOLOGY | 1536 | 1% | 1% | 3 |
| 5 | CURRENT OPINION IN BIOTECHNOLOGY | 1315 | 1% | 0% | 7 |
| 6 | JOURNAL OF MICROBIOLOGICAL METHODS | 1283 | 2% | 0% | 10 |
| 7 | MICROBIAL CELL FACTORIES | 1151 | 1% | 0% | 5 |
| 8 | COMPUTATIONAL BIOLOGY AND CHEMISTRY | 1139 | 1% | 0% | 4 |
| 9 | BMC GENOMICS | 1087 | 3% | 0% | 13 |
| 10 | ANNUAL REVIEW OF GENOMICS AND HUMAN GENETICS | 777 | 0% | 1% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ESSENTIAL GENES | 610103 | 9% | 23% | 43 | Search ESSENTIAL+GENES | Search ESSENTIAL+GENES |
| 2 | TN SEQ | 236324 | 1% | 54% | 7 | Search TN+SEQ | Search TN+SEQ |
| 3 | SUBTRACTIVE GENOMICS | 225715 | 1% | 60% | 6 | Search SUBTRACTIVE+GENOMICS | Search SUBTRACTIVE+GENOMICS |
| 4 | MINIMAL GENOME | 188088 | 2% | 33% | 9 | Search MINIMAL+GENOME | Search MINIMAL+GENOME |
| 5 | ALLELIC REPLACEMENT MUTAGENESIS | 125400 | 0% | 100% | 2 | Search ALLELIC+REPLACEMENT+MUTAGENESIS | Search ALLELIC+REPLACEMENT+MUTAGENESIS |
| 6 | COMPUTATIONAL COMPARATIVE GENOMICS | 125400 | 0% | 100% | 2 | Search COMPUTATIONAL+COMPARATIVE+GENOMICS | Search COMPUTATIONAL+COMPARATIVE+GENOMICS |
| 7 | ESSENTIAL GENE PREDICTION | 125400 | 0% | 100% | 2 | Search ESSENTIAL+GENE+PREDICTION | Search ESSENTIAL+GENE+PREDICTION |
| 8 | ESSENTIAL GENOME | 125400 | 0% | 100% | 2 | Search ESSENTIAL+GENOME | Search ESSENTIAL+GENOME |
| 9 | IN SILICO DRUG TARGETS | 125400 | 0% | 100% | 2 | Search IN+SILICO+DRUG+TARGETS | Search IN+SILICO+DRUG+TARGETS |
| 10 | KAAS SERVER | 125400 | 0% | 100% | 2 | Search KAAS+SERVER | Search KAAS+SERVER |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LUO, H , LIN, Y , GAO, F , ZHANG, CT , ZHANG, R , (2014) DEG 10, AN UPDATE OF THE DATABASE OF ESSENTIAL GENES THAT INCLUDES BOTH PROTEIN-CODING GENES AND NONCODING GENOMIC ELEMENTS.NUCLEIC ACIDS RESEARCH. VOL. 42. ISSUE D1. P. D574-D580 | 36 | 49% | 77 |
| 2 | LU, Y , DENG, JY , CARSON, MB , LU, H , LU, LJ , (2014) COMPUTATIONAL METHODS FOR THE PREDICTION OF MICROBIAL ESSENTIAL GENES.CURRENT BIOINFORMATICS. VOL. 9. ISSUE 2. P. 89 -101 | 38 | 42% | 1 |
| 3 | ZHANG, R , LIN, Y , (2009) DEG 5.0, A DATABASE OF ESSENTIAL GENES IN BOTH PROKARYOTES AND EUKARYOTES.NUCLEIC ACIDS RESEARCH. VOL. 37. ISSUE . P. D455 -D458 | 21 | 54% | 182 |
| 4 | DENG, JY , SU, SC , LIN, XD , HASSETT, DJ , LU, LJ , (2013) A STATISTICAL FRAMEWORK FOR IMPROVING GENOMIC ANNOTATIONS OF PROKARYOTIC ESSENTIAL GENES.PLOS ONE. VOL. 8. ISSUE 3. P. - | 24 | 56% | 6 |
| 5 | GAO, F , ZHANG, RR , (2011) ENZYMES ARE ENRICHED IN BACTERIAL ESSENTIAL GENES.PLOS ONE. VOL. 6. ISSUE 6. P. - | 22 | 59% | 6 |
| 6 | WEI, W , NING, LW , YE, YN , GUO, FB , (2013) GEPTOP: A GENE ESSENTIALITY PREDICTION TOOL FOR SEQUENCED BACTERIAL GENOMES BASED ON ORTHOLOGY AND PHYLOGENY.PLOS ONE. VOL. 8. ISSUE 8. P. - | 24 | 51% | 5 |
| 7 | LIN, Y , ZHANG, RR , (2011) PUTATIVE ESSENTIAL AND CORE-ESSENTIAL GENES IN MYCOPLASMA GENOMES.SCIENTIFIC REPORTS. VOL. 1. ISSUE . P. - | 19 | 66% | 12 |
| 8 | RICKE, SC , MANDAL, RK , KWON, YM , (2016) TRANSPOSON SEQUENCING: METHODS AND EXPANDING APPLICATIONS.APPLIED MICROBIOLOGY AND BIOTECHNOLOGY. VOL. 100. ISSUE 1. P. 31 -43 | 28 | 32% | 4 |
| 9 | CHAO, MC , ABEL, S , DAVIS, BM , WALDOR, MK , (2016) THE DESIGN AND ANALYSIS OF TRANSPOSON INSERTION SEQUENCING EXPERIMENTS.NATURE REVIEWS MICROBIOLOGY. VOL. 14. ISSUE 2. P. 119 -128 | 24 | 40% | 2 |
| 10 | CHENG, J , XU, Z , WU, WW , ZHAO, L , LI, XC , LIU, YL , TAO, SH , (2014) TRAINING SET SELECTION FOR THE PREDICTION OF ESSENTIAL GENES.PLOS ONE. VOL. 9. ISSUE 1. P. - | 23 | 46% | 3 |
Classes with closest relation at Level 1 |