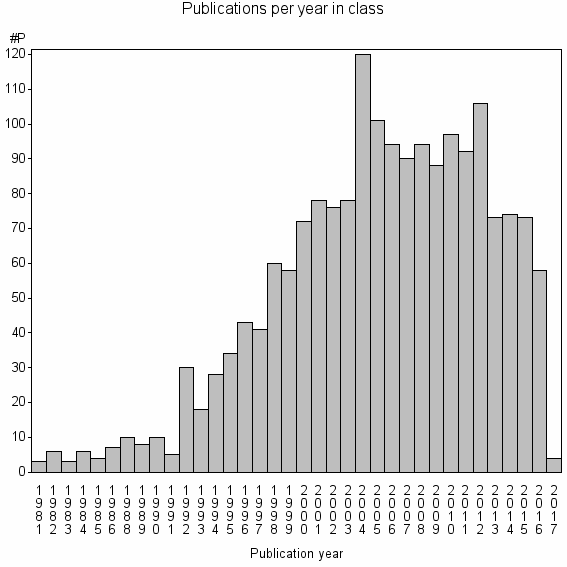
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 4131 | 1842 | 50.0 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 365 | 2 | PROTEIN STRUCTURE PREDICTION//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PSEUDO AMINO ACID COMPOSITION | 17331 |
| 4131 | 1 | TRANSMEMBRANE HELIX//TRANSMEMBRANE HELICES//MEMBRANE PROTEIN | 1842 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | TRANSMEMBRANE HELIX | authKW | 439352 | 3% | 43% | 62 |
| 2 | TRANSMEMBRANE HELICES | authKW | 319097 | 2% | 51% | 38 |
| 3 | MEMBRANE PROTEIN | authKW | 279449 | 15% | 6% | 284 |
| 4 | HELIX HELIX INTERACTION | authKW | 215803 | 1% | 52% | 25 |
| 5 | MEMBRANE PROTEIN FOLDING | authKW | 197324 | 2% | 37% | 32 |
| 6 | TOPOLOGY PREDICTION | authKW | 191602 | 1% | 68% | 17 |
| 7 | HYDROPHOBIC MISMATCH | authKW | 133358 | 1% | 31% | 26 |
| 8 | TRANSMEMBRANE SEGMENT | authKW | 120084 | 1% | 32% | 23 |
| 9 | TRANSMEMBRANE HELIX PREDICTION | authKW | 117869 | 0% | 89% | 8 |
| 10 | TOXCAT | authKW | 116708 | 1% | 54% | 13 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biophysics | 12427 | 30% | 0% | 553 |
| 2 | Biochemistry & Molecular Biology | 12360 | 66% | 0% | 1222 |
| 3 | Mathematical & Computational Biology | 2907 | 8% | 0% | 139 |
| 4 | Biochemical Research Methods | 1557 | 10% | 0% | 193 |
| 5 | Biotechnology & Applied Microbiology | 378 | 7% | 0% | 138 |
| 6 | Cell Biology | 343 | 9% | 0% | 165 |
| 7 | Computer Science, Interdisciplinary Applications | 253 | 4% | 0% | 70 |
| 8 | Biology | 40 | 2% | 0% | 38 |
| 9 | Chemistry, Physical | 9 | 5% | 0% | 94 |
| 10 | Chemistry, Multidisciplinary | 8 | 5% | 0% | 88 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | STOCKHOLM BIOINFORMAT | 93413 | 2% | 17% | 34 |
| 2 | MOL STRUCT FUNCT | 87955 | 2% | 17% | 31 |
| 3 | SCI BIO OURCES PROGRAM | 82878 | 0% | 100% | 5 |
| 4 | BIOMEMBRANE | 53903 | 3% | 6% | 52 |
| 5 | LEHRSTUHL CHEM BIOPOLYMERE | 51722 | 1% | 20% | 16 |
| 6 | ELECT INFORMAT SYST ENGN | 41767 | 1% | 23% | 11 |
| 7 | BIOCHEM MEMBRANES | 37600 | 1% | 12% | 19 |
| 8 | PROGRAM MACROMOL STRUCT | 36837 | 1% | 17% | 13 |
| 9 | GENOM PROTEMOL BIOL | 33151 | 0% | 100% | 2 |
| 10 | SWEDEN BIOINFORMAT INFRASTRUCT LIFE SCI BILS | 33151 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOPHYSICAL JOURNAL | 16321 | 8% | 1% | 139 |
| 2 | BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES | 15671 | 4% | 1% | 77 |
| 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 14911 | 4% | 1% | 76 |
| 4 | JOURNAL OF MOLECULAR BIOLOGY | 13622 | 8% | 1% | 139 |
| 5 | PROTEIN SCIENCE | 11080 | 4% | 1% | 65 |
| 6 | BIOCHEMISTRY | 10957 | 10% | 0% | 189 |
| 7 | BIOINFORMATICS | 6684 | 4% | 1% | 66 |
| 8 | CURRENT OPINION IN STRUCTURAL BIOLOGY | 6507 | 2% | 1% | 30 |
| 9 | BIOPOLYMERS | 4876 | 2% | 1% | 44 |
| 10 | COMPUTATIONAL BIOLOGY AND CHEMISTRY | 4233 | 1% | 2% | 15 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TRANSMEMBRANE HELIX | 439352 | 3% | 43% | 62 | Search TRANSMEMBRANE+HELIX | Search TRANSMEMBRANE+HELIX |
| 2 | TRANSMEMBRANE HELICES | 319097 | 2% | 51% | 38 | Search TRANSMEMBRANE+HELICES | Search TRANSMEMBRANE+HELICES |
| 3 | MEMBRANE PROTEIN | 279449 | 15% | 6% | 284 | Search MEMBRANE+PROTEIN | Search MEMBRANE+PROTEIN |
| 4 | HELIX HELIX INTERACTION | 215803 | 1% | 52% | 25 | Search HELIX+HELIX+INTERACTION | Search HELIX+HELIX+INTERACTION |
| 5 | MEMBRANE PROTEIN FOLDING | 197324 | 2% | 37% | 32 | Search MEMBRANE+PROTEIN+FOLDING | Search MEMBRANE+PROTEIN+FOLDING |
| 6 | TOPOLOGY PREDICTION | 191602 | 1% | 68% | 17 | Search TOPOLOGY+PREDICTION | Search TOPOLOGY+PREDICTION |
| 7 | HYDROPHOBIC MISMATCH | 133358 | 1% | 31% | 26 | Search HYDROPHOBIC+MISMATCH | Search HYDROPHOBIC+MISMATCH |
| 8 | TRANSMEMBRANE SEGMENT | 120084 | 1% | 32% | 23 | Search TRANSMEMBRANE+SEGMENT | Search TRANSMEMBRANE+SEGMENT |
| 9 | TRANSMEMBRANE HELIX PREDICTION | 117869 | 0% | 89% | 8 | Search TRANSMEMBRANE+HELIX+PREDICTION | Search TRANSMEMBRANE+HELIX+PREDICTION |
| 10 | TOXCAT | 116708 | 1% | 54% | 13 | Search TOXCAT | Search TOXCAT |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LI, E , WIMLEY, WC , HRISTOVA, K , (2012) TRANSMEMBRANE HELIX DIMERIZATION: BEYOND THE SEARCH FOR SEQUENCE MOTIFS.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1818. ISSUE 2. P. 183-193 | 100 | 82% | 54 |
| 2 | MACKENZIE, KR , (2006) FOLDING AND STABILITY OF ALPHA-HELICAL INTEGRAL MEMBRANE PROTEINS.CHEMICAL REVIEWS. VOL. 106. ISSUE 5. P. 1931 -1977 | 242 | 38% | 133 |
| 3 | STEINDORF, D , SCHNEIDER, D , (2017) IN VIVO SELECTION OF HETEROTYPICALLY INTERACTING TRANSMEMBRANE HELICES: COMPLEMENTARY HELIX SURFACES, RATHER THAN CONSERVED INTERACTION MOTIFS, DRIVE FORMATION OF TRANSMEMBRANE HETERO-DIMERS.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1859. ISSUE 2. P. 245 -256 | 83 | 93% | 0 |
| 4 | BORDAG, N , KELLER, S , (2010) ALPHA-HELICAL TRANSMEMBRANE PEPTIDES: A "DIVIDE AND CONQUER" APPROACH TO MEMBRANE PROTEINS.CHEMISTRY AND PHYSICS OF LIPIDS. VOL. 163. ISSUE 1. P. 1-26 | 165 | 39% | 44 |
| 5 | LIANG, J , NAVEED, H , JIMENEZ-MORALES, D , ADAMIAN, L , LIN, MS , (2012) COMPUTATIONAL STUDIES OF MEMBRANE PROTEINS: MODELS AND PREDICTIONS FOR BIOLOGICAL UNDERSTANDING.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1818. ISSUE 4. P. 927 -941 | 105 | 59% | 12 |
| 6 | TUSNADY, GE , SIMON, I , (2010) TOPOLOGY PREDICTION OF HELICAL TRANSMEMBRANE PROTEINS: HOW FAR HAVE WE REACHED?.CURRENT PROTEIN & PEPTIDE SCIENCE. VOL. 11. ISSUE 7. P. 550 -561 | 83 | 72% | 14 |
| 7 | RATH, A , DEBER, CM , (2012) PROTEIN STRUCTURE IN MEMBRANE DOMAINS.ANNUAL REVIEW OF BIOPHYSICS, VOL 41. VOL. 41. ISSUE . P. 135-155 | 68 | 82% | 11 |
| 8 | SENES, A , ENGEL, DE , DEGRADO, WF , (2004) FOLDING OF HELICAL MEMBRANE PROTEINS: THE ROLE OF POLAR, GXXXG-LIKE AND PROLINE MOTIFS.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 14. ISSUE 4. P. 465 -479 | 83 | 62% | 271 |
| 9 | LANGOSCH, D , ARKIN, IT , (2009) INTERACTION AND CONFORMATIONAL DYNAMICS OF MEMBRANE-SPANNING PROTEIN HELICES.PROTEIN SCIENCE. VOL. 18. ISSUE 7. P. 1343-1358 | 105 | 50% | 61 |
| 10 | CYMER, F , VEERAPPAN, A , SCHNEIDER, D , (2012) TRANSMEMBRANE HELIX-HELIX INTERACTIONS ARE MODULATED BY THE SEQUENCE CONTEXT AND BY LIPID BILAYER PROPERTIES.BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES. VOL. 1818. ISSUE 4. P. 963 -973 | 65 | 72% | 37 |
Classes with closest relation at Level 1 |