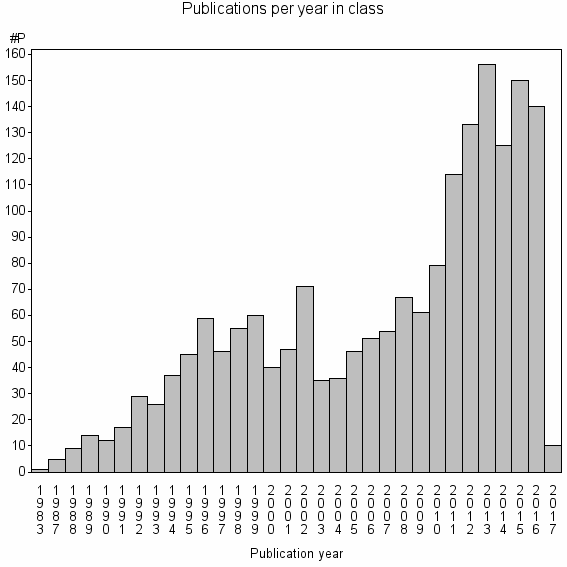
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 4196 | 1830 | 52.7 | 89% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 405 | 3 | BACTERIORHODOPSIN//G PROTEIN COUPLED RECEPTOR//G PROTEIN | 29641 |
| 705 | 2 | G PROTEIN COUPLED RECEPTOR//GPCR//RHODOPSIN | 12950 |
| 4196 | 1 | G PROTEIN COUPLED RECEPTOR//GPCR//MEMBRANE PROT STRUCT FUNCT UNIT | 1830 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | G PROTEIN COUPLED RECEPTOR | authKW | 230198 | 14% | 5% | 257 |
| 2 | GPCR | authKW | 164598 | 8% | 7% | 141 |
| 3 | MEMBRANE PROT STRUCT FUNCT UNIT | address | 150152 | 1% | 75% | 12 |
| 4 | CONSTITUTIVE ACTIVITY | authKW | 129015 | 2% | 18% | 42 |
| 5 | MED COMPUTAC | address | 124053 | 2% | 25% | 30 |
| 6 | IONIC LOCK | authKW | 90833 | 0% | 78% | 7 |
| 7 | GPCRS | authKW | 90730 | 2% | 14% | 40 |
| 8 | UNITAT BIOESTADIST | address | 78980 | 2% | 16% | 30 |
| 9 | RECEPTOR ACTIVATION | authKW | 78935 | 1% | 18% | 27 |
| 10 | GPCR ACTIVATION | authKW | 71179 | 0% | 53% | 8 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 7061 | 52% | 0% | 945 |
| 2 | Chemistry, Medicinal | 3714 | 14% | 0% | 254 |
| 3 | Pharmacology & Pharmacy | 2896 | 25% | 0% | 454 |
| 4 | Biophysics | 2123 | 13% | 0% | 241 |
| 5 | Computer Science, Interdisciplinary Applications | 1386 | 8% | 0% | 146 |
| 6 | Biochemical Research Methods | 517 | 7% | 0% | 119 |
| 7 | Cell Biology | 463 | 10% | 0% | 184 |
| 8 | Mathematical & Computational Biology | 320 | 3% | 0% | 50 |
| 9 | Computer Science, Information Systems | 163 | 3% | 0% | 56 |
| 10 | Chemistry, Multidisciplinary | 149 | 9% | 0% | 164 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MEMBRANE PROT STRUCT FUNCT UNIT | 150152 | 1% | 75% | 12 |
| 2 | MED COMPUTAC | 124053 | 2% | 25% | 30 |
| 3 | UNITAT BIOESTADIST | 78980 | 2% | 16% | 30 |
| 4 | OULU BIO | 38132 | 0% | 57% | 4 |
| 5 | MED CHEM NEUROENGN | 35581 | 0% | 27% | 8 |
| 6 | PHARMACEUT INTELLIGENCE PLATFORM | 33368 | 0% | 100% | 2 |
| 7 | PSYCHIAT MOLEC NEUROPHARMACOL SECT | 33368 | 0% | 100% | 2 |
| 8 | SECT THEORET RECEPTOR PHARMACOL | 33368 | 0% | 100% | 2 |
| 9 | SYNTH CHEM TECHNOL PHARMACEUT SUBST COMP M | 33368 | 0% | 100% | 2 |
| 10 | VAN ANDEL CAS RECEPTOR | 33368 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR PHARMACOLOGY | 24955 | 7% | 1% | 121 |
| 2 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | 16167 | 3% | 2% | 55 |
| 3 | RECEPTORS & CHANNELS | 13572 | 1% | 5% | 16 |
| 4 | TRENDS IN PHARMACOLOGICAL SCIENCES | 11727 | 3% | 1% | 49 |
| 5 | JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN | 8444 | 2% | 2% | 30 |
| 6 | JOURNAL OF RECEPTOR AND SIGNAL TRANSDUCTION RESEARCH | 3999 | 0% | 3% | 8 |
| 7 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 3520 | 2% | 1% | 37 |
| 8 | JOURNAL OF MOLECULAR MODELING | 2208 | 1% | 1% | 22 |
| 9 | MOLECULAR INFORMATICS | 2159 | 0% | 2% | 8 |
| 10 | JOURNAL OF BIOLOGICAL CHEMISTRY | 2066 | 8% | 0% | 143 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | G PROTEIN COUPLED RECEPTOR | 230198 | 14% | 5% | 257 | Search G+PROTEIN+COUPLED+RECEPTOR | Search G+PROTEIN+COUPLED+RECEPTOR |
| 2 | GPCR | 164598 | 8% | 7% | 141 | Search GPCR | Search GPCR |
| 3 | CONSTITUTIVE ACTIVITY | 129015 | 2% | 18% | 42 | Search CONSTITUTIVE+ACTIVITY | Search CONSTITUTIVE+ACTIVITY |
| 4 | IONIC LOCK | 90833 | 0% | 78% | 7 | Search IONIC+LOCK | Search IONIC+LOCK |
| 5 | GPCRS | 90730 | 2% | 14% | 40 | Search GPCRS | Search GPCRS |
| 6 | RECEPTOR ACTIVATION | 78935 | 1% | 18% | 27 | Search RECEPTOR+ACTIVATION | Search RECEPTOR+ACTIVATION |
| 7 | GPCR ACTIVATION | 71179 | 0% | 53% | 8 | Search GPCR+ACTIVATION | Search GPCR+ACTIVATION |
| 8 | CONFORMATIONAL THERMOSTABILIZATION | 69516 | 0% | 83% | 5 | Search CONFORMATIONAL+THERMOSTABILIZATION | Search CONFORMATIONAL+THERMOSTABILIZATION |
| 9 | GPCR STRUCTURE | 66729 | 0% | 50% | 8 | Search GPCR+STRUCTURE | Search GPCR+STRUCTURE |
| 10 | ROTAMER TOGGLE SWITCH | 53388 | 0% | 80% | 4 | Search ROTAMER+TOGGLE+SWITCH | Search ROTAMER+TOGGLE+SWITCH |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | VENKATAKRISHNAN, AJ , DEUPI, X , LEBON, G , TATE, CG , SCHERTLER, GF , BABU, MM , (2013) MOLECULAR SIGNATURES OF G-PROTEIN-COUPLED RECEPTORS.NATURE. VOL. 494. ISSUE 7436. P. 185-194 | 62 | 65% | 412 |
| 2 | KATRITCH, V , CHEREZOV, V , STEVENS, RC , (2013) STRUCTURE-FUNCTION OF THE G PROTEIN-COUPLED RECEPTOR SUPERFAMILY.ANNUAL REVIEW OF PHARMACOLOGY AND TOXICOLOGY, VOL 53, 2013. VOL. 53. ISSUE . P. 531-556 | 71 | 62% | 289 |
| 3 | FANELLI, F , DE BENEDETTI, PG , (2011) UPDATE 1 OF: COMPUTATIONAL MODELING APPROACHES TO STRUCTURE-FUNCTION ANALYSIS OF G PROTEIN-COUPLED RECEPTORS.CHEMICAL REVIEWS. VOL. 111. ISSUE . P. PR438 -PR535 | 320 | 27% | 38 |
| 4 | CAVASOTTO, CN , PALOMBA, D , (2015) EXPANDING THE HORIZONS OF G PROTEIN-COUPLED RECEPTOR STRUCTURE-BASED LIGAND DISCOVERY AND OPTIMIZATION USING HOMOLOGY MODELS.CHEMICAL COMMUNICATIONS. VOL. 51. ISSUE 71. P. 13576 -13594 | 137 | 53% | 9 |
| 5 | KONTOYIANNI, M , (2016) G PROTEIN COUPLED RECEPTORS AND STRUCTURE-BASED ADVANCES.CURRENT TOPICS IN MEDICINAL CHEMISTRY. VOL. 16. ISSUE 13. P. 1489 -1505 | 111 | 55% | 1 |
| 6 | RODRIGUEZ, D , RANGANATHAN, A , CARLSSON, J , (2015) DISCOVERY OF GPCR LIGANDS BY MOLECULAR DOCKING SCREENING: NOVEL OPPORTUNITIES PROVIDED BY CRYSTAL STRUCTURES.CURRENT TOPICS IN MEDICINAL CHEMISTRY. VOL. 15. ISSUE 24. P. 2484 -2503 | 95 | 62% | 8 |
| 7 | ANDREWS, SP , BROWN, GA , CHRISTOPHER, JA , (2014) STRUCTURE-BASED AND FRAGMENT-BASED GPCR DRUG DISCOVERY.CHEMMEDCHEM. VOL. 9. ISSUE 2. P. 256-275 | 87 | 61% | 27 |
| 8 | RODRIGUEZ, D , GUTIERREZ-DE-TERAN, H , (2013) COMPUTATIONAL APPROACHES FOR LIGAND DISCOVERY AND DESIGN IN CLASS-A G PROTEIN-COUPLED RECEPTORS.CURRENT PHARMACEUTICAL DESIGN. VOL. 19. ISSUE 12. P. 2216 -2236 | 111 | 54% | 7 |
| 9 | LATORRACA, NR , VENKATAKRISHNAN, AJ , DROR, RO , (2017) GPCR DYNAMICS: STRUCTURES IN MOTION.CHEMICAL REVIEWS. VOL. 117. ISSUE 1. P. 139 -155 | 72 | 47% | 1 |
| 10 | STEVENS, RC , CHEREZOV, V , KATRITCH, V , ABAGYAN, R , KUHN, P , ROSEN, H , WUTHRICH, K , (2013) THE GPCR NETWORK: A LARGE-SCALE COLLABORATION TO DETERMINE HUMAN GPCR STRUCTURE AND FUNCTION.NATURE REVIEWS DRUG DISCOVERY. VOL. 12. ISSUE 1. P. 25 -34 | 49 | 83% | 88 |
Classes with closest relation at Level 1 |