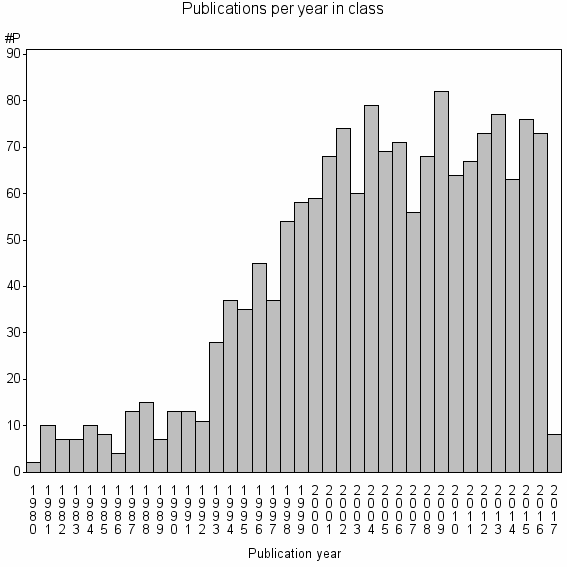
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 5584 | 1601 | 47.8 | 88% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 153 | 3 | BIOCHEMISTRY & MOLECULAR BIOLOGY//RIBOSOME//RNA | 62593 |
| 1297 | 2 | RIBOSOME//TRANSLATION TERMINATION//TMRNA | 8599 |
| 5584 | 1 | ATP DEPENDENT PROTEASE//CLPB//FTSH | 1601 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ATP DEPENDENT PROTEASE | authKW | 1043614 | 4% | 79% | 69 |
| 2 | CLPB | authKW | 711578 | 3% | 75% | 50 |
| 3 | FTSH | authKW | 611399 | 3% | 70% | 46 |
| 4 | LON PROTEASE | authKW | 611213 | 3% | 64% | 50 |
| 5 | LON | authKW | 382953 | 2% | 57% | 35 |
| 6 | ATP DEPENDENT PROTEOLYSIS | authKW | 325014 | 2% | 61% | 28 |
| 7 | CLP PROTEASE | authKW | 322281 | 2% | 65% | 26 |
| 8 | ATP DEPENDENT PROTEASES | authKW | 305128 | 1% | 80% | 20 |
| 9 | CLPA | authKW | 268184 | 1% | 94% | 15 |
| 10 | HSP104 | authKW | 256308 | 2% | 40% | 34 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 10631 | 66% | 0% | 1057 |
| 2 | Cell Biology | 3711 | 26% | 0% | 411 |
| 3 | Microbiology | 3270 | 20% | 0% | 317 |
| 4 | Biophysics | 2019 | 14% | 0% | 219 |
| 5 | Plant Sciences | 493 | 9% | 0% | 147 |
| 6 | Genetics & Heredity | 91 | 5% | 0% | 74 |
| 7 | Biochemical Research Methods | 58 | 3% | 0% | 48 |
| 8 | Biology | 13 | 2% | 0% | 25 |
| 9 | Crystallography | 12 | 1% | 0% | 23 |
| 10 | Biotechnology & Applied Microbiology | 9 | 3% | 0% | 42 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | AVIRU | 76284 | 0% | 100% | 4 |
| 2 | EXIST TRANSFER | 57213 | 0% | 100% | 3 |
| 3 | MOL MIKROBIOL | 38142 | 0% | 100% | 2 |
| 4 | UNITE BIOCHIM MICROBIENNE | 36596 | 1% | 16% | 12 |
| 5 | ZENTRUM MOL BIOL HEIDELBERG | 27110 | 0% | 18% | 8 |
| 6 | BIOCHEM MOL BIOL IBMB | 25427 | 0% | 67% | 2 |
| 7 | NRL PROT BIOCHEM | 21451 | 0% | 38% | 3 |
| 8 | BIOELE OCHEM NANOTECHNOL GRP | 19071 | 0% | 100% | 1 |
| 9 | BIOL 68523 | 19071 | 0% | 100% | 1 |
| 10 | BIOL CHEMCLUSTER EXCELLENCE MACROMOL COMPLEX | 19071 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF STRUCTURAL BIOLOGY | 11593 | 3% | 1% | 46 |
| 2 | JOURNAL OF BACTERIOLOGY | 6963 | 7% | 0% | 111 |
| 3 | MOLECULAR MICROBIOLOGY | 5893 | 4% | 1% | 61 |
| 4 | JOURNAL OF BIOLOGICAL CHEMISTRY | 3495 | 11% | 0% | 171 |
| 5 | MOLECULAR CELL | 2894 | 2% | 1% | 29 |
| 6 | NATURE STRUCTURAL & MOLECULAR BIOLOGY | 2520 | 1% | 1% | 17 |
| 7 | JOURNAL OF MOLECULAR BIOLOGY | 1735 | 3% | 0% | 47 |
| 8 | STRUCTURE | 1423 | 1% | 0% | 17 |
| 9 | RESEARCH IN MICROBIOLOGY | 1394 | 1% | 1% | 14 |
| 10 | PLANT CELL | 1375 | 1% | 0% | 21 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ATP DEPENDENT PROTEASE | 1043614 | 4% | 79% | 69 | Search ATP+DEPENDENT+PROTEASE | Search ATP+DEPENDENT+PROTEASE |
| 2 | CLPB | 711578 | 3% | 75% | 50 | Search CLPB | Search CLPB |
| 3 | FTSH | 611399 | 3% | 70% | 46 | Search FTSH | Search FTSH |
| 4 | LON PROTEASE | 611213 | 3% | 64% | 50 | Search LON+PROTEASE | Search LON+PROTEASE |
| 5 | LON | 382953 | 2% | 57% | 35 | Search LON | Search LON |
| 6 | ATP DEPENDENT PROTEOLYSIS | 325014 | 2% | 61% | 28 | Search ATP+DEPENDENT+PROTEOLYSIS | Search ATP+DEPENDENT+PROTEOLYSIS |
| 7 | CLP PROTEASE | 322281 | 2% | 65% | 26 | Search CLP+PROTEASE | Search CLP+PROTEASE |
| 8 | ATP DEPENDENT PROTEASES | 305128 | 1% | 80% | 20 | Search ATP+DEPENDENT+PROTEASES | Search ATP+DEPENDENT+PROTEASES |
| 9 | CLPA | 268184 | 1% | 94% | 15 | Search CLPA | Search CLPA |
| 10 | HSP104 | 256308 | 2% | 40% | 34 | Search HSP104 | Search HSP104 |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SAUER, RT , BAKER, TA , (2011) AAA+ PROTEASES: ATP-FUELED MACHINES OF PROTEIN DESTRUCTION.ANNUAL REVIEW OF BIOCHEMISTRY, VOL 80. VOL. 80. ISSUE . P. 587-612 | 113 | 71% | 206 |
| 2 | BAKER, TA , SAUER, RT , (2012) CLPXP, AN ATP-POWERED UNFOLDING AND PROTEIN-DEGRADATION MACHINE.BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH. VOL. 1823. ISSUE 1. P. 15-28 | 90 | 83% | 93 |
| 3 | OLIVARES, AO , BAKER, TA , SAUER, RT , (2016) MECHANISTIC INSIGHTS INTO BACTERIAL AAA PLUS PROTEASES AND PROTEIN-REMODELLING MACHINES.NATURE REVIEWS MICROBIOLOGY. VOL. 14. ISSUE 1. P. 33 -44 | 74 | 69% | 12 |
| 4 | BITTNER, LM , ARENDS, J , NARBERHAUS, F , (2016) ATP-DEPENDENT PROTEASES IN BACTERIA.BIOPOLYMERS. VOL. 105. ISSUE 8. P. 505 -517 | 107 | 78% | 0 |
| 5 | HODSON, S , MARSHALL, JJT , BURSTON, SG , (2012) MAPPING THE ROAD TO RECOVERY: THE CLPB/HSP104 MOLECULAR CHAPERONE.JOURNAL OF STRUCTURAL BIOLOGY. VOL. 179. ISSUE 2. P. 161-171 | 79 | 87% | 16 |
| 6 | VENKATESH, S , LEE, J , SINGH, K , LEE, I , SUZUKI, CK , (2012) MULTITASKING IN THE MITOCHONDRION BY THE ATP-DEPENDENT LON PROTEASE.BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH. VOL. 1823. ISSUE 1. P. 56-66 | 84 | 65% | 58 |
| 7 | DESANTIS, ME , SHORTER, J , (2012) THE ELUSIVE MIDDLE DOMAIN OF HSP104 AND CLPB: LOCATION AND FUNCTION.BIOCHIMICA ET BIOPHYSICA ACTA-MOLECULAR CELL RESEARCH. VOL. 1823. ISSUE 1. P. 29-39 | 89 | 68% | 24 |
| 8 | DOYLE, SM , WICKNER, S , (2009) HSP104 AND CLPB: PROTEIN DISAGGREGATING MACHINES.TRENDS IN BIOCHEMICAL SCIENCES. VOL. 34. ISSUE 1. P. 40 -48 | 64 | 80% | 128 |
| 9 | ITO, K , AKIYAMA, Y , (2005) CELLULAR FUNCTIONS, MECHANISM OF ACTION, AND REGULATION OF FTSH PROTEASE.ANNUAL REVIEW OF MICROBIOLOGY. VOL. 59. ISSUE . P. 211 -231 | 74 | 76% | 206 |
| 10 | LEE, I , SUZUKI, CK , (2008) FUNCTIONAL MECHANICS OF THE ATP-DEPENDENT LON PROTEASE-LESSONS FROM ENDOGENOUS PROTEIN AND SYNTHETIC PEPTIDE SUBSTRATES.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1784. ISSUE 5. P. 727-735 | 75 | 82% | 44 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 16075 | HRCA//CIRCE//SIGMA32 |
| 2 | 11831 | SUP35//YEAST PRION//URE2P |
| 3 | 15103 | HTRA1//HTRA//HTRA2 |
| 4 | 18779 | TMRNA//TRANS TRANSLATION//SMPB |
| 5 | 31973 | PUPYLATION//PROKARYOTIC UBIQUITIN LIKE PROTEIN//PAFA |
| 6 | 22758 | ARGINYLATION//JOHANSON BLIZZARD SYNDROME//N END RULE |
| 7 | 6152 | DNAJ//DNAK//GRPE |
| 8 | 24380 | BACTERIOPHAGE LAMBDA DEVELOPMENT//LAMBDA PLASMIDS//COLIPHAGE LAMBDA |
| 9 | 1429 | PROTEASOME//MULTICATALYTIC PROTEINASE//IMMUNOPROTEASOME |
| 10 | 11129 | D1 PROTEIN//PHOTOINHIBITION//PHOTOSYSTEM II |