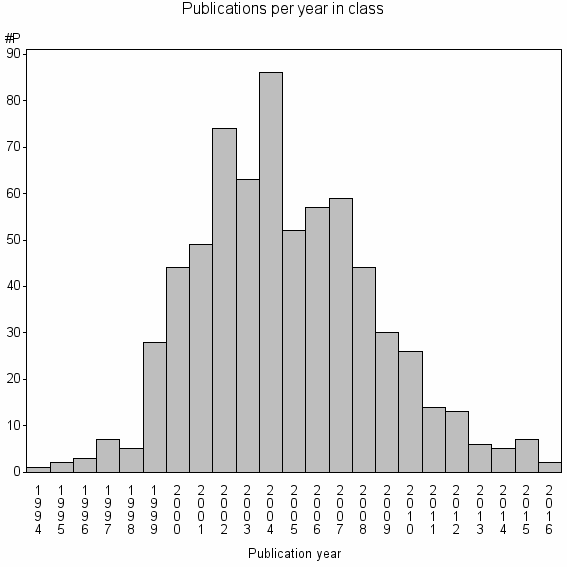
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 16034 | 677 | 35.4 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 2924 | 2 | SAGE//SERIAL ANALYSIS OF GENE EXPRESSION//DIFFERENTIAL DISPLAY | 2737 |
| 16034 | 1 | SERIAL ANALYSIS OF GENE EXPRESSION//SAGE//GLGI | 677 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | SERIAL ANALYSIS OF GENE EXPRESSION | authKW | 561554 | 6% | 30% | 41 |
| 2 | SAGE | authKW | 511656 | 12% | 14% | 79 |
| 3 | GLGI | authKW | 180410 | 1% | 100% | 4 |
| 4 | SERIAL ANALYSIS OF GENE EXPRESSION SAGE | authKW | 169367 | 2% | 29% | 13 |
| 5 | TAISHO FUNCT GENOM | address | 145479 | 1% | 32% | 10 |
| 6 | SAGE TECHNIQUE | authKW | 135308 | 0% | 100% | 3 |
| 7 | ENH | address | 93948 | 1% | 21% | 10 |
| 8 | ADAPTER TAGGED COMPETITIVE PCR | authKW | 90205 | 0% | 100% | 2 |
| 9 | ADAPTOR TAGGED COMPETITIVE POLYMERASE CHAIN REACTION | authKW | 90205 | 0% | 100% | 2 |
| 10 | DIGITAL GENE EXPRESSION DISPLAYER | authKW | 90205 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 3009 | 29% | 0% | 195 |
| 2 | Genetics & Heredity | 2797 | 27% | 0% | 185 |
| 3 | Mathematical & Computational Biology | 1392 | 9% | 0% | 58 |
| 4 | Biochemical Research Methods | 831 | 12% | 0% | 84 |
| 5 | Biochemistry & Molecular Biology | 771 | 31% | 0% | 207 |
| 6 | Ophthalmology | 194 | 5% | 0% | 35 |
| 7 | Cell Biology | 158 | 10% | 0% | 66 |
| 8 | Information Science & Library Science | 107 | 2% | 0% | 15 |
| 9 | Oncology | 82 | 8% | 0% | 53 |
| 10 | Multidisciplinary Sciences | 35 | 1% | 0% | 10 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | TAISHO FUNCT GENOM | 145479 | 1% | 32% | 10 |
| 2 | ENH | 93948 | 1% | 21% | 10 |
| 3 | EQUIPE SIGNALISAT IDENT CELLULAI | 90205 | 0% | 100% | 2 |
| 4 | FUNCT GENOM ANAT PHYSIOL | 90205 | 0% | 100% | 2 |
| 5 | IMAGE CONSORTIUM | 67651 | 0% | 50% | 3 |
| 6 | MOL ENDOCRINOL ONCOL ANAT PHYSIO | 60135 | 0% | 67% | 2 |
| 7 | AGR ENVIRONM SCI GENOM IL | 45103 | 0% | 100% | 1 |
| 8 | ANAT PHYSIOLCHUL | 45103 | 0% | 100% | 1 |
| 9 | ANAT PHYSIOLFUNCT GENOM | 45103 | 0% | 100% | 1 |
| 10 | AXIS TEAM PROJECT | 45103 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BMC GENOMICS | 6414 | 5% | 0% | 37 |
| 2 | BIOTECHFORUM ( BTF ) | 6268 | 3% | 1% | 19 |
| 3 | GENOMICS | 4242 | 4% | 0% | 28 |
| 4 | SCIENTIST | 4157 | 2% | 1% | 15 |
| 5 | PHYSIOLOGICAL GENOMICS | 4092 | 2% | 1% | 13 |
| 6 | BMC BIOINFORMATICS | 4027 | 4% | 0% | 26 |
| 7 | MOLECULAR VISION | 3065 | 2% | 0% | 15 |
| 8 | CURRENT PHARMACEUTICAL BIOTECHNOLOGY | 2113 | 1% | 1% | 8 |
| 9 | BIOINFORMATICS | 2017 | 3% | 0% | 22 |
| 10 | GENOME BIOLOGY | 1811 | 2% | 0% | 11 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ANISIMOV, SV , (2008) SERIAL ANALYSIS OF GENE EXPRESSION (SAGE): 13 YEARS OF APPLICATION IN RESEARCH.CURRENT PHARMACEUTICAL BIOTECHNOLOGY. VOL. 9. ISSUE 5. P. 338-350 | 86 | 67% | 37 |
| 2 | WANG, SM , (2007) UNDERSTANDING SAGE DATA.TRENDS IN GENETICS. VOL. 23. ISSUE 1. P. 42 -50 | 33 | 70% | 49 |
| 3 | WANG, HY , ZHENG, HR , AZUAJE, F , (2008) CLUSTERING-BASED APPROACHES TO SAGE DATA MINING.BIODATA MINING. VOL. 1. ISSUE . P. - | 31 | 76% | 5 |
| 4 | TUTEJA, R , TUTEJA, N , (2004) SERIAL ANALYSIS OF GENE EXPRESSION (SAGE): UNRAVELING THE BIOINFORMATICS TOOLS.BIOESSAYS. VOL. 26. ISSUE 8. P. 916-922 | 29 | 78% | 16 |
| 5 | HU, M , POLYAK, K , (2006) SERIAL ANALYSIS OF GENE EXPRESSION.NATURE PROTOCOLS. VOL. 1. ISSUE 4. P. 1743 -1760 | 39 | 53% | 14 |
| 6 | XU, WJ , WANG, ZX , QIAO, ZD , (2009) MODIFIED PCR METHODS FOR 3' END AMPLIFICATION FROM SERIAL ANALYSIS OF GENE EXPRESSION (SAGE) TAGS.FEBS JOURNAL. VOL. 276. ISSUE 10. P. 2657 -2668 | 28 | 67% | 1 |
| 7 | BARTLETT, J , (2001) TECHNOLOGY EVALUATION: SAGE, GENZYME MOLECULAR ONCOLOGY.CURRENT OPINION IN MOLECULAR THERAPEUTICS. VOL. 3. ISSUE 1. P. 85 -96 | 40 | 51% | 14 |
| 8 | VEGA-SANCHEZ, ME , GOWDA, M , WANG, GL , (2007) TAG-BASED APPROACHES FOR DEEP TRANSCRIPTOME ANALYSIS IN PLANTS.PLANT SCIENCE. VOL. 173. ISSUE 4. P. 371-380 | 32 | 53% | 7 |
| 9 | PATINO, WD , MIAN, OY , HWANG, PM , (2002) SERIAL ANALYSIS OF GENE EXPRESSION - TECHNICAL CONSIDERATIONS AND APPLICATIONS TO CARDIOVASCULAR BIOLOGY.CIRCULATION RESEARCH. VOL. 91. ISSUE 7. P. 565 -569 | 27 | 69% | 30 |
| 10 | POROYKO, V , HEJLEK, LG , SPOLLEN, WG , SPRINGER, GK , NGUYEN, HT , SHARP, RE , BOHNERT, HJ , (2005) THE MAIZE ROOT TRANSCRIPTOME BY SERIAL ANALYSIS OF GENE EXPRESSION.PLANT PHYSIOLOGY. VOL. 138. ISSUE 3. P. 1700 -1710 | 24 | 69% | 44 |
Classes with closest relation at Level 1 |