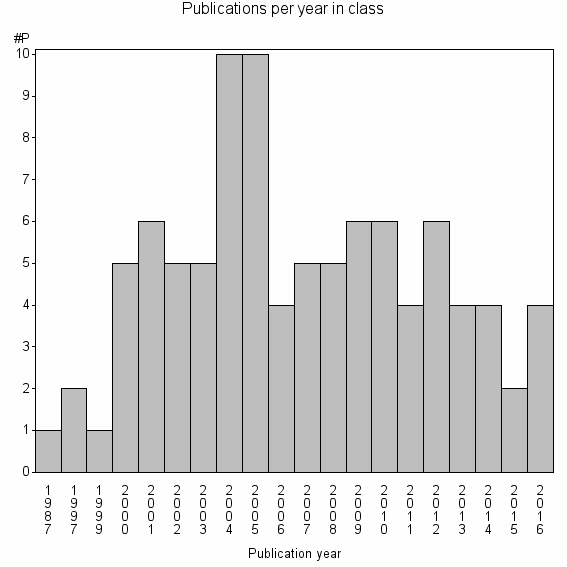
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 35565 | 95 | 27.6 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 2924 | 2 | SAGE//SERIAL ANALYSIS OF GENE EXPRESSION//DIFFERENTIAL DISPLAY | 2737 |
| 35565 | 1 | MKIAA//ATAD3//ATAD3A | 95 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | MKIAA | authKW | 2057134 | 8% | 80% | 8 |
| 2 | ATAD3 | authKW | 1653055 | 6% | 86% | 6 |
| 3 | ATAD3A | authKW | 857138 | 4% | 67% | 4 |
| 4 | LARGE PROTEINS | authKW | 788946 | 9% | 27% | 9 |
| 5 | INTEGRATED DATABASE SYST BIOL TEAM | address | 723211 | 3% | 75% | 3 |
| 6 | LONG CDNA | authKW | 723211 | 3% | 75% | 3 |
| 7 | SINGLE PASS SEQUENCE | authKW | 723211 | 3% | 75% | 3 |
| 8 | CDNA SEQUENCING | authKW | 677394 | 15% | 15% | 14 |
| 9 | ATPASE FAMILY AAA DOMAIN CONTAINING 3A | authKW | 642855 | 2% | 100% | 2 |
| 10 | MIHANA KU | address | 642855 | 2% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 781 | 38% | 0% | 36 |
| 2 | Biochemistry & Molecular Biology | 193 | 39% | 0% | 37 |
| 3 | Biochemical Research Methods | 64 | 9% | 0% | 9 |
| 4 | Mathematical & Computational Biology | 43 | 4% | 0% | 4 |
| 5 | Cell Biology | 34 | 12% | 0% | 11 |
| 6 | Biotechnology & Applied Microbiology | 7 | 5% | 0% | 5 |
| 7 | Biology | 7 | 3% | 0% | 3 |
| 8 | Biophysics | 7 | 4% | 0% | 4 |
| 9 | Medicine, Research & Experimental | 6 | 4% | 0% | 4 |
| 10 | Evolutionary Biology | 1 | 1% | 0% | 1 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | INTEGRATED DATABASE SYST BIOL TEAM | 723211 | 3% | 75% | 3 |
| 2 | MIHANA KU | 642855 | 2% | 100% | 2 |
| 3 | INTEGRATED DATABASE GRP | 342850 | 4% | 27% | 4 |
| 4 | ANDRE LWOFFUMR 935 | 321428 | 1% | 100% | 1 |
| 5 | CLIN COOPERATION GRP MOL ONCOL | 321428 | 1% | 100% | 1 |
| 6 | EQUIPE BIOMICS MITOSTEM | 321428 | 1% | 100% | 1 |
| 7 | ESTEAM PARIS SUD STEM CELL CORE ILINSERM | 321428 | 1% | 100% | 1 |
| 8 | GENOME KNOWLEDGEBASE | 321428 | 1% | 100% | 1 |
| 9 | GENOME TASK GRP | 321428 | 1% | 100% | 1 |
| 10 | GENOMES FUNCT EXPLORAT | 321428 | 1% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DNA RESEARCH | 202117 | 21% | 3% | 20 |
| 2 | NUCLEIC ACIDS RESEARCH | 2641 | 19% | 0% | 18 |
| 3 | HUMAN GENOMICS | 2509 | 1% | 1% | 1 |
| 4 | BIOMOLECULAR ENGINEERING | 957 | 1% | 0% | 1 |
| 5 | INTERNATIONAL JOURNAL OF DIABETES IN DEVELOPING COUNTRIES | 744 | 1% | 0% | 1 |
| 6 | JOURNAL OF BIOENERGETICS AND BIOMEMBRANES | 672 | 2% | 0% | 2 |
| 7 | GENE | 591 | 6% | 0% | 6 |
| 8 | CYTOLOGY AND GENETICS | 571 | 1% | 0% | 1 |
| 9 | INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE | 525 | 3% | 0% | 3 |
| 10 | ACTA BIOLOGICA CRACOVIENSIA SERIES BOTANICA | 476 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MKIAA | 2057134 | 8% | 80% | 8 | Search MKIAA | Search MKIAA |
| 2 | ATAD3 | 1653055 | 6% | 86% | 6 | Search ATAD3 | Search ATAD3 |
| 3 | ATAD3A | 857138 | 4% | 67% | 4 | Search ATAD3A | Search ATAD3A |
| 4 | LARGE PROTEINS | 788946 | 9% | 27% | 9 | Search LARGE+PROTEINS | Search LARGE+PROTEINS |
| 5 | LONG CDNA | 723211 | 3% | 75% | 3 | Search LONG+CDNA | Search LONG+CDNA |
| 6 | SINGLE PASS SEQUENCE | 723211 | 3% | 75% | 3 | Search SINGLE+PASS+SEQUENCE | Search SINGLE+PASS+SEQUENCE |
| 7 | CDNA SEQUENCING | 677394 | 15% | 15% | 14 | Search CDNA+SEQUENCING | Search CDNA+SEQUENCING |
| 8 | ATPASE FAMILY AAA DOMAIN CONTAINING 3A | 642855 | 2% | 100% | 2 | Search ATPASE+FAMILY+AAA+DOMAIN+CONTAINING+3A | Search ATPASE+FAMILY+AAA+DOMAIN+CONTAINING+3A |
| 9 | TOB3 | 642855 | 2% | 100% | 2 | Search TOB3 | Search TOB3 |
| 10 | PROTEIN CODING REGION | 365251 | 5% | 23% | 5 | Search PROTEIN+CODING+REGION | Search PROTEIN+CODING+REGION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LI, SJ , LAMARCHE, F , CHARTON, R , DELPHIN, C , GIRES, O , HUBSTENBERGER, A , SCHLATTNER, U , ROUSSEAU, D , (2014) EXPRESSION ANALYSIS OF ATAD3 ISOFORMS IN RODENT AND HUMAN CELL LINES AND TISSUES.GENE. VOL. 535. ISSUE 1. P. 60-69 | 11 | 48% | 4 |
| 2 | KOGA, H , (2006) ESTABLISHMENT OF THE PLATFORM FOR REVERSE CHEMICAL GENETICS TARGETING NOVEL PROTEIN-PROTEIN INTERACTIONS.MOLECULAR BIOSYSTEMS. VOL. 2. ISSUE 3-4. P. 159-164 | 13 | 46% | 1 |
| 3 | OKAZAKI, N , F-KIKUNO, R , OHARA, R , INAMOTO, S , KOSEKI, H , HIRAOKA, S , SAGA, Y , SEINO, S , NISHIMURA, M , KAISHO, T , ET AL (2004) PREDICTION OF THE CODING SEQUENCES OF MOUSE HOMOLOGUES OF KIAA GENE: IV. THE COMPLETE NUCLEOTIDE SEQUENCES OF 500 MOUSE KIAA-HOMOLOGOUS CDNAS IDENTIFIED BY SCREENING OF TERMINAL SEQUENCES OF CDNA CLONES RANDOMLY SAMPLED FROM SIZE-FRACTIONATED LIBRARIES.DNA RESEARCH. VOL. 11. ISSUE 3. P. 205-218 | 12 | 50% | 8 |
| 4 | KIKUNO, R , NAGASE, T , NAKAYAMA, M , KOGA, H , OKAZAKI, N , NAKAJIMA, D , OHARA, O , (2004) HUGE: A DATABASE FOR HUMAN KIAA PROTEINS, A 2004 UPDATE INTEGRATING HUGEPPI AND ROUGE.NUCLEIC ACIDS RESEARCH. VOL. 32. ISSUE . P. D502 -D504 | 8 | 57% | 40 |
| 5 | OKAZAKI, N , KIKUNO, R , OHARA, R , INAMOTO, S , KOSEKI, H , HIRAOKA, S , SAGA, Y , KITAMURA, H , NAKAGAWA, T , NAGASE, T , ET AL (2004) PREDICTION OF THE CODING SEQUENCES OF MOUSE HOMOLOGUES OF FLJ GENES: THE COMPLETE NUCLEOTIDE SEQUENCES OF 110 MOUSE FLJ-HOMOLOGOUS CDNAS IDENTIFIED BY SCREENING OF TERMINAL SEQUENCES OF CDNA CLONES RANDOMLY SAMPLED FROM SIZE-FRACTIONATED LIBRARIES.DNA RESEARCH. VOL. 11. ISSUE 2. P. 127-135 | 9 | 47% | 2 |
| 6 | IMANISHI, T , NAGAI, Y , HABARA, T , YAMASAKI, C , TAKEDA, J , MIKAMI, S , BANDO, Y , TOJO, H , NISHIMURA, T , (2013) FULL-LENGTH TRANSCRIPTOME-BASED H-INVDB THROWS A NEW LIGHT ON CHROMOSOME-CENTRIC PROTEOMICS.JOURNAL OF PROTEOME RESEARCH. VOL. 12. ISSUE 1. P. 62 -66 | 6 | 55% | 3 |
| 7 | SHIMADA, K , NAGANO, M , KAWAI, M , KOGA, H , (2005) INFLUENCES OF AMINO ACID FEATURES OF GLUTATHIONE S-TRANSFERASE FUSION PROTEINS ON THEIR SOLUBILITY.PROTEOMICS. VOL. 5. ISSUE 15. P. 3859-3863 | 9 | 45% | 5 |
| 8 | TAKEDA, J , YAMASAKI, C , MURAKAMI, K , NAGAI, Y , SERA, M , HARA, Y , OBI, N , HABARA, T , GOJOBORI, T , IMANISHI, T , (2013) H-INVDB IN 2013: AN OMICS STUDY PLATFORM FOR HUMAN FUNCTIONAL GENE AND TRANSCRIPT DISCOVERY.NUCLEIC ACIDS RESEARCH. VOL. 41. ISSUE D1. P. D915 -D919 | 9 | 33% | 15 |
| 9 | LI, SJ , ROUSSEAU, D , (2012) ATAD3, A VITAL MEMBRANE BOUND MITOCHONDRIAL ATPASE INVOLVED IN TUMOR PROGRESSION.JOURNAL OF BIOENERGETICS AND BIOMEMBRANES. VOL. 44. ISSUE 1. P. 189-197 | 9 | 33% | 11 |
| 10 | VAN DEN ECKER, D , HOFFMANN, M , MUTING, G , MAGLIONI, S , HEREBIAN, D , MAYATEPEK, E , VENTURA, N , DISTELMAIER, F , (2015) CAENORHABDITIS ELEGANS ATAD-3 MODULATES MITOCHONDRIAL IRON AND HEME HOMEOSTASIS.BIOCHEMICAL AND BIOPHYSICAL RESEARCH COMMUNICATIONS. VOL. 467. ISSUE 2. P. 389 -394 | 10 | 26% | 0 |
Classes with closest relation at Level 1 |