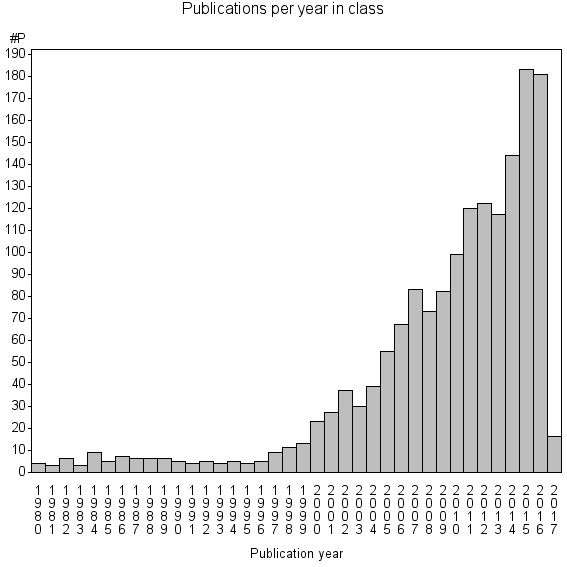
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 5481 | 1618 | 49.1 | 91% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 71 | 3 | GENETICS & HEREDITY//CHROMATIN//CELL BIOLOGY | 83302 |
| 343 | 2 | EZH2//CHROMATIN//POLYCOMB | 17754 |
| 5481 | 1 | ARGININE METHYLATION//PROTEIN ARGININE METHYLTRANSFERASE//PRMT | 1618 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ARGININE METHYLATION | authKW | 1216711 | 6% | 65% | 99 |
| 2 | PROTEIN ARGININE METHYLTRANSFERASE | authKW | 753761 | 3% | 74% | 54 |
| 3 | PRMT | authKW | 592001 | 2% | 78% | 40 |
| 4 | PROTEIN ARGININE METHYLATION | authKW | 537579 | 2% | 81% | 35 |
| 5 | PRMT1 | authKW | 507375 | 3% | 61% | 44 |
| 6 | CARM1 | authKW | 507297 | 2% | 79% | 34 |
| 7 | SMYD3 | authKW | 386550 | 2% | 79% | 26 |
| 8 | LYSINE METHYLATION | authKW | 290224 | 2% | 42% | 37 |
| 9 | PRMT5 | authKW | 283368 | 2% | 52% | 29 |
| 10 | G9A | authKW | 259655 | 2% | 40% | 34 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 8461 | 59% | 0% | 958 |
| 2 | Cell Biology | 4557 | 28% | 0% | 454 |
| 3 | Biophysics | 762 | 9% | 0% | 143 |
| 4 | Biochemical Research Methods | 661 | 8% | 0% | 123 |
| 5 | Chemistry, Medicinal | 513 | 6% | 0% | 97 |
| 6 | Developmental Biology | 384 | 3% | 0% | 56 |
| 7 | Oncology | 349 | 10% | 0% | 156 |
| 8 | Genetics & Heredity | 306 | 7% | 0% | 115 |
| 9 | Cell & Tissue Engineering | 31 | 0% | 0% | 8 |
| 10 | Chemistry, Analytical | 9 | 3% | 0% | 43 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | TERRY FOX MOL ONCOL GRP | 73402 | 1% | 16% | 24 |
| 2 | STRUCT GENOM CONSORTIUM | 55670 | 3% | 6% | 50 |
| 3 | PROGRAM MOL BASIS DIS | 42457 | 0% | 75% | 3 |
| 4 | DECS STRUCT CHEM | 37741 | 0% | 100% | 2 |
| 5 | FRE2618 | 37741 | 0% | 100% | 2 |
| 6 | HORT PLANT PHYSIOL BIOCHEM | 37741 | 0% | 100% | 2 |
| 7 | PARASITE CELL BIOL GRP | 37741 | 0% | 100% | 2 |
| 8 | TRI TRAINING PROGRAM CHEM BIOL | 37725 | 1% | 20% | 10 |
| 9 | BLOOMFIELD AGING | 34820 | 1% | 8% | 22 |
| 10 | EXPT INFECT DIS | 31320 | 1% | 12% | 14 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ACS CHEMICAL BIOLOGY | 3836 | 1% | 1% | 21 |
| 2 | MOLECULAR CELL | 3716 | 2% | 1% | 33 |
| 3 | JOURNAL OF BIOLOGICAL CHEMISTRY | 3005 | 10% | 0% | 160 |
| 4 | GENES & DEVELOPMENT | 2086 | 2% | 0% | 29 |
| 5 | EPIGENETICS & CHROMATIN | 1623 | 0% | 2% | 5 |
| 6 | MOLECULAR & CELLULAR PROTEOMICS | 1539 | 1% | 1% | 16 |
| 7 | HRVATSKI FILMSKI LJETOPIS | 1256 | 0% | 7% | 1 |
| 8 | EPIGENOMICS | 1011 | 0% | 1% | 5 |
| 9 | CHEMBIOCHEM | 928 | 1% | 0% | 15 |
| 10 | MEDCHEMCOMM | 880 | 0% | 1% | 8 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ARGININE METHYLATION | 1216711 | 6% | 65% | 99 | Search ARGININE+METHYLATION | Search ARGININE+METHYLATION |
| 2 | PROTEIN ARGININE METHYLTRANSFERASE | 753761 | 3% | 74% | 54 | Search PROTEIN+ARGININE+METHYLTRANSFERASE | Search PROTEIN+ARGININE+METHYLTRANSFERASE |
| 3 | PRMT | 592001 | 2% | 78% | 40 | Search PRMT | Search PRMT |
| 4 | PROTEIN ARGININE METHYLATION | 537579 | 2% | 81% | 35 | Search PROTEIN+ARGININE+METHYLATION | Search PROTEIN+ARGININE+METHYLATION |
| 5 | PRMT1 | 507375 | 3% | 61% | 44 | Search PRMT1 | Search PRMT1 |
| 6 | CARM1 | 507297 | 2% | 79% | 34 | Search CARM1 | Search CARM1 |
| 7 | SMYD3 | 386550 | 2% | 79% | 26 | Search SMYD3 | Search SMYD3 |
| 8 | LYSINE METHYLATION | 290224 | 2% | 42% | 37 | Search LYSINE+METHYLATION | Search LYSINE+METHYLATION |
| 9 | PRMT5 | 283368 | 2% | 52% | 29 | Search PRMT5 | Search PRMT5 |
| 10 | G9A | 259655 | 2% | 40% | 34 | Search G9A | Search G9A |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HU, H , QIAN, K , HO, MC , ZHENG, YG , (2016) SMALL MOLECULE INHIBITORS OF PROTEIN ARGININE METHYLTRANSFERASES.EXPERT OPINION ON INVESTIGATIONAL DRUGS. VOL. 25. ISSUE 3. P. 335 -358 | 166 | 77% | 6 |
| 2 | YANG, YZ , BEDFORD, MT , (2013) PROTEIN ARGININE METHYLTRANSFERASES AND CANCER.NATURE REVIEWS CANCER. VOL. 13. ISSUE 1. P. 37 -50 | 127 | 67% | 157 |
| 3 | MORALES, Y , CACERES, T , MAY, K , HEVEL, JM , (2016) BIOCHEMISTRY AND REGULATION OF THE PROTEIN ARGININE METHYLTRANSFERASES (PRMTS).ARCHIVES OF BIOCHEMISTRY AND BIOPHYSICS. VOL. 590. ISSUE . P. 138 -152 | 106 | 69% | 9 |
| 4 | BEDFORD, MT , CLARKE, SG , (2009) PROTEIN ARGININE METHYLATION IN MAMMALS: WHO, WHAT, AND WHY.MOLECULAR CELL. VOL. 33. ISSUE 1. P. 1-13 | 85 | 73% | 558 |
| 5 | KUHN, P , XU, W , (2009) PROTEIN ARGININE METHYLTRANSFERASES: NUCLEAR RECEPTOR COREGULATORS AND BEYOND.REGULATORY MECHANISMS IN TRANSCRIPTIONAL SIGNALING. VOL. 87. ISSUE . P. 299-342 | 145 | 81% | 7 |
| 6 | WEI, H , MUNDADE, R , LANGE, KC , LU, T , (2014) PROTEIN ARGININE METHYLATION OF NON-HISTONE PROTEINS AND ITS ROLE IN DISEASES.CELL CYCLE. VOL. 13. ISSUE 1. P. 32-41 | 90 | 88% | 30 |
| 7 | MORETTIN, A , BALDWIN, RM , COTE, J , (2015) ARGININE METHYLTRANSFERASES AS NOVEL THERAPEUTIC TARGETS FOR BREAST CANCER.MUTAGENESIS. VOL. 30. ISSUE 2. P. 177 -189 | 128 | 64% | 4 |
| 8 | WOLF, SS , (2009) THE PROTEIN ARGININE METHYLTRANSFERASE FAMILY: AN UPDATE ABOUT FUNCTION, NEW PERSPECTIVES AND THE PHYSIOLOGICAL ROLE IN HUMANS.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 66. ISSUE 13. P. 2109 -2121 | 94 | 85% | 84 |
| 9 | KANISKAN, HU , KONZE, KD , JIN, J , (2015) SELECTIVE INHIBITORS OF PROTEIN METHYLTRANSFERASES.JOURNAL OF MEDICINAL CHEMISTRY. VOL. 58. ISSUE 4. P. 1596 -1629 | 126 | 47% | 25 |
| 10 | STOPA, N , KREBS, JE , SHECHTER, D , (2015) THE PRMT5 ARGININE METHYLTRANSFERASE: MANY ROLES IN DEVELOPMENT, CANCER AND BEYOND.CELLULAR AND MOLECULAR LIFE SCIENCES. VOL. 72. ISSUE 11. P. 2041 -2059 | 100 | 59% | 17 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 12023 | SET8//HISTONE//PR SET7 |
| 2 | 8082 | HISTONE DEMETHYLASE//LSD1//HISTONE DEMETHYLATION |
| 3 | 14125 | USP22//SET1//UBIQUITIN SPECIFIC PROTEASE 22 |
| 4 | 27625 | METABOLIC MEMORY//IL RISK ASSESSMENT INTERVENT STUDIES//HYPERGLYCEMIC MEMORY |
| 5 | 24691 | BTG2//BTG1//TOB |
| 6 | 1710 | EZH2//POLYCOMB//BMI 1 |
| 7 | 7575 | HP1//POSITION EFFECT VARIEGATION//HETEROCHROMATIN |
| 8 | 9000 | STEROID HORMONES SECT//AIB1//SRC 3 |
| 9 | 20432 | KINASE SPECIFIC//PTM CODE//CELLULAR MOL LOG TEAM |
| 10 | 16735 | EVI1//EVI 1//RIZ1 |