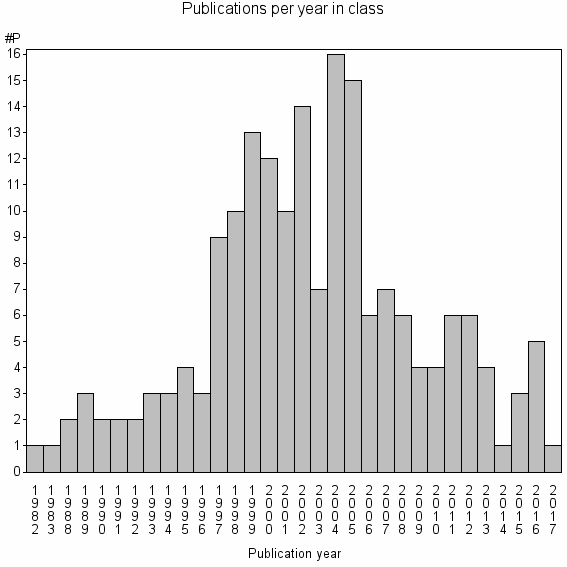
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 29487 | 185 | 52.2 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 304 | 3 | P53//RETINOBLASTOMA//MDM2 | 40663 |
| 936 | 2 | P53//MDM2//P73 | 10991 |
| 29487 | 1 | PINK EYED UNSTABLE//RADIAT ON AB MOL CELLULAR RADIAT BIOL//3 5 EXONUCLEASE | 185 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PINK EYED UNSTABLE | authKW | 330113 | 1% | 100% | 2 |
| 2 | RADIAT ON AB MOL CELLULAR RADIAT BIOL | address | 247582 | 2% | 50% | 3 |
| 3 | 3 5 EXONUCLEASE | authKW | 222811 | 5% | 15% | 9 |
| 4 | ARE RNA | authKW | 220074 | 1% | 67% | 2 |
| 5 | INTRACHROMOSOMAL RECOMBINATION | authKW | 212206 | 3% | 21% | 6 |
| 6 | 3 INDOLE | authKW | 165057 | 1% | 100% | 1 |
| 7 | 3 TERMINAL MISMATCHED DNA | authKW | 165057 | 1% | 100% | 1 |
| 8 | ALTERNATIVELY SPLICED P53 | authKW | 165057 | 1% | 100% | 1 |
| 9 | C FOS AND P53 | authKW | 165057 | 1% | 100% | 1 |
| 10 | CARCINOGEN INDUCED DELETION | authKW | 165057 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 2003 | 43% | 0% | 80 |
| 2 | Toxicology | 1086 | 23% | 0% | 43 |
| 3 | Cell Biology | 792 | 34% | 0% | 63 |
| 4 | Oncology | 766 | 34% | 0% | 63 |
| 5 | Biochemistry & Molecular Biology | 478 | 43% | 0% | 80 |
| 6 | Biotechnology & Applied Microbiology | 93 | 11% | 0% | 20 |
| 7 | Biophysics | 20 | 5% | 0% | 9 |
| 8 | Biology | 9 | 3% | 0% | 5 |
| 9 | Medicine, Research & Experimental | 5 | 3% | 0% | 6 |
| 10 | Virology | 4 | 2% | 0% | 3 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | RADIAT ON AB MOL CELLULAR RADIAT BIOL | 247582 | 2% | 50% | 3 |
| 2 | CNRSUNITE MIXTE RECH 217 | 165057 | 1% | 100% | 1 |
| 3 | DIRECT SCI VIVANTDEP RADIOBIOL RADIOPATHOL | 165057 | 1% | 100% | 1 |
| 4 | FDN LELOIR INVEST BIOQUIM BUENO AI | 165057 | 1% | 100% | 1 |
| 5 | LINEBERGER COMPREHENS CANC 119 | 165057 | 1% | 100% | 1 |
| 6 | MED CURRICULUM TOXICOL | 165057 | 1% | 100% | 1 |
| 7 | MOL MED COLOGNE CMMC SYST BIOL AGEING COLOG | 165057 | 1% | 100% | 1 |
| 8 | OSP SEREGNO | 165057 | 1% | 100% | 1 |
| 9 | RADIAT ON AB MOL CELLLULAR RADIAT BIOL | 165057 | 1% | 100% | 1 |
| 10 | RADIAT ON AB MOL CELLULARE RADIAT BIOL | 165057 | 1% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ONCOGENE | 7706 | 16% | 0% | 29 |
| 2 | MUTATION RESEARCH-DNA REPAIR | 5415 | 2% | 1% | 4 |
| 3 | MUTATION RESEARCH-FUNDAMENTAL AND MOLECULAR MECHANISMS OF MUTAGENESIS | 3423 | 5% | 0% | 9 |
| 4 | MUTAGENESIS | 2625 | 3% | 0% | 6 |
| 5 | CARCINOGENESIS | 2396 | 7% | 0% | 13 |
| 6 | DNA REPAIR | 2022 | 3% | 0% | 5 |
| 7 | IN VITRO & MOLECULAR TOXICOLOGY-A JOURNAL OF BASIC AND APPLIED RESEARCH | 1682 | 1% | 1% | 1 |
| 8 | MUTATION RESEARCH-GENETIC TOXICOLOGY AND ENVIRONMENTAL MUTAGENESIS | 792 | 2% | 0% | 4 |
| 9 | MUTATION RESEARCH | 724 | 3% | 0% | 6 |
| 10 | MUTATION RESEARCH-GENETIC TOXICOLOGY | 571 | 1% | 0% | 1 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PINK EYED UNSTABLE | 330113 | 1% | 100% | 2 | Search PINK+EYED+UNSTABLE | Search PINK+EYED+UNSTABLE |
| 2 | 3 5 EXONUCLEASE | 222811 | 5% | 15% | 9 | Search 3+5+EXONUCLEASE | Search 3+5+EXONUCLEASE |
| 3 | ARE RNA | 220074 | 1% | 67% | 2 | Search ARE+RNA | Search ARE+RNA |
| 4 | INTRACHROMOSOMAL RECOMBINATION | 212206 | 3% | 21% | 6 | Search INTRACHROMOSOMAL+RECOMBINATION | Search INTRACHROMOSOMAL+RECOMBINATION |
| 5 | 3 INDOLE | 165057 | 1% | 100% | 1 | Search 3+INDOLE | Search 3+INDOLE |
| 6 | 3 TERMINAL MISMATCHED DNA | 165057 | 1% | 100% | 1 | Search 3+TERMINAL+MISMATCHED+DNA | Search 3+TERMINAL+MISMATCHED+DNA |
| 7 | ALTERNATIVELY SPLICED P53 | 165057 | 1% | 100% | 1 | Search ALTERNATIVELY+SPLICED+P53 | Search ALTERNATIVELY+SPLICED+P53 |
| 8 | C FOS AND P53 | 165057 | 1% | 100% | 1 | Search C+FOS+AND+P53 | Search C+FOS+AND+P53 |
| 9 | CARCINOGEN INDUCED DELETION | 165057 | 1% | 100% | 1 | Search CARCINOGEN+INDUCED+DELETION | Search CARCINOGEN+INDUCED+DELETION |
| 10 | CELL CYCLE RADIATION | 165057 | 1% | 100% | 1 | Search CELL+CYCLE+RADIATION | Search CELL+CYCLE+RADIATION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GATZ, SA , WIESMULLER, L , (2006) P53 IN RECOMBINATION AND REPAIR.CELL DEATH AND DIFFERENTIATION. VOL. 13. ISSUE 6. P. 1003-1016 | 50 | 31% | 115 |
| 2 | SAINTIGNY, Y , BERTRAND, P , LOPEZ, BS , (2005) DIRECT ROLE OF P53 ON HOMOLOGOUS RECOMBINATION.M S-MEDECINE SCIENCES. VOL. 21. ISSUE 1. P. 43-48 | 22 | 59% | 2 |
| 3 | RIECKMANN, T , KRIEGS, M , NITSCH, L , HOFFER, K , ROHALY, G , KOCHER, S , PETERSEN, C , DIKOMEY, E , DORNREITER, I , DAHM-DAPHI, J , (2013) P53 MODULATES HOMOLOGOUS RECOMBINATION AT I-SCEI-INDUCED DOUBLE-STRAND BREAKS THROUGH CELL-CYCLE REGULATION.ONCOGENE. VOL. 32. ISSUE 8. P. 968-975 | 18 | 51% | 10 |
| 4 | SENGUPTA, S , HARRIS, CC , (2005) P53: TRAFFIC COP AT THE CROSSROADS OF DNA REPAIR AND RECOMBINATION.NATURE REVIEWS MOLECULAR CELL BIOLOGY. VOL. 6. ISSUE 1. P. 44-55 | 31 | 22% | 286 |
| 5 | KEIMLING, M , WIESMULLER, L , (2009) DNA DOUBLE-STRAND BREAK REPAIR ACTIVITIES IN MAMMARY EPITHELIAL CELLS--INFLUENCE OF ENDOGENOUS P53 VARIANTS.CARCINOGENESIS. VOL. 30. ISSUE 7. P. 1260 -1268 | 20 | 40% | 12 |
| 6 | BERTRAND, P , SAINTIGNY, Y , LOPEZ, BS , (2004) P53 ' S DOUBLE LIFE: TRANSACTIVATION-INDEPENDENT REPRESSION OF HOMOLOGOUS RECOMBINATION.TRENDS IN GENETICS. VOL. 20. ISSUE 6. P. 235-243 | 23 | 34% | 92 |
| 7 | SIRBU, BM , LACHMAYER, SJ , WULFING, V , MARTEN, LM , CLARKSON, KE , LEE, LW , GHEORGHIU, L , ZOU, L , POWELL, SN , DAHM-DAPHI, J , ET AL (2011) ATR-P53 RESTRICTS HOMOLOGOUS RECOMBINATION IN RESPONSE TO REPLICATIVE STRESS BUT DOES NOT LIMIT DNA INTERSTRAND CROSSLINK REPAIR IN LUNG CANCER CELLS.PLOS ONE. VOL. 6. ISSUE 8. P. - | 22 | 32% | 7 |
| 8 | WIKTOR-BROWN, DM , SUKUP-JACKSON, MR , FAKHRALDEEN, SA , HENDRICKS, CA , ENGELWARD, BP , (2011) P53 NULL FLUORESCENT YELLOW DIRECT REPEAT (FYDR) MICE HAVE NORMAL LEVELS OF HOMOLOGOUS RECOMBINATION.DNA REPAIR. VOL. 10. ISSUE 12. P. 1294-1299 | 22 | 31% | 5 |
| 9 | HAMPPA, S , KIESSLINGA, T , BUECHLEA, K , MANSILLA, SF , THOMALEC, J , RALL, M , AHN, J , POSPIECH, H , GOTTIFREDI, V , WIESMULLER, L , (2016) DNA DAMAGE TOLERANCE PATHWAY INVOLVING DNA POLYMERASE IOTA AND THE TUMOR SUPPRESSOR P53 REGULATES DNA REPLICATION FORK PROGRESSION.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 113. ISSUE 30. P. E4311 -E4319 | 20 | 27% | 2 |
| 10 | AKYUZ, N , BOEHDEN, GS , SUSSE, S , RIMEK, A , PREUSS, U , SCHEIDTMANN, KH , WIESMULLER, L , (2002) DNA SUBSTRATE DEPENDENCE OF P53-MEDIATED REGULATION OF DOUBLE-STRAND BREAK REPAIR.MOLECULAR AND CELLULAR BIOLOGY. VOL. 22. ISSUE 17. P. 6306-6317 | 19 | 36% | 97 |
Classes with closest relation at Level 1 |