Protease engineering and protease substrate profiling
Proteases play a crucial role in all organisms by regulating a myriad of biological processes through their ability to catalyze the hydrolysis of peptide bonds. Recombinant proteases are today important tools in biotechnological and biomedical research and are also used as therapeutics. Engineering of proteases with tailor-made properties is thus important.
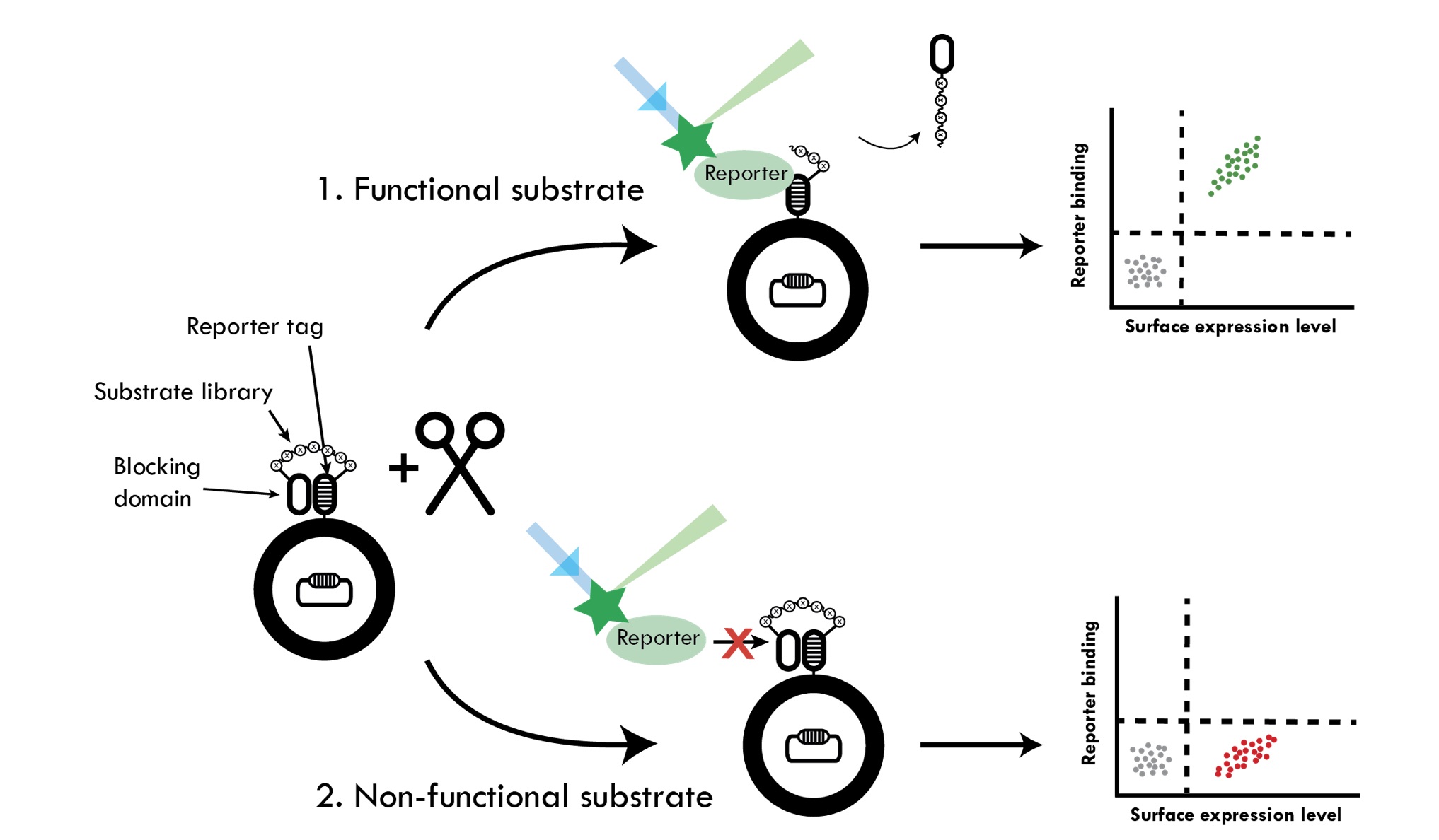
We are developing several different high-throughput methods for quantitative assessment of relative proteolytic activity. The methods are based on either expression of substrate libraries on the surface of bacteria followed by incubation with respective protease or co-expression of substrates and proteases in the cytoplasm. An important property of proteases is to discriminate among potential substrates. The former method is thus developed for substrate profiling of proteases. The substrates are displayed between two so-called anti-idiotypic affinity domains on bacteria. The first domain functions as a reporter tag and has affinity for a labeled reporter protein, whereas the second domain blocks the reporter tag from interacting with the reporter protein. Site-specific proteolysis of the substrate results in release of the blocking domain, enabling the reporter tag to bind the labeled reporter protein. Proteolysis is therefore reflected in reporter binding. The latter method involves an aggregation-prone peptide that is fused to a fluorescent protein via a linker containing the respective protease substrate sequence. Cleavage of the substrate sequence in the reporter rescues the fluorescent protein from aggregation and results in increased whole-cell fluorescence correlating to proteolytic activity, which can be monitored. In both methods we are sorting the bacterial libraries using flow cytometry, which allows for screening of millions of variants. The huge libraries that can be constructed in bacteria and the extreme throughput of modern flow cytometers make these methods very powerful.
Research funded by: the Swedish Research Council