Nyhetsflöde
Logga in till din kurswebb
Du är inte inloggad på KTH så innehållet är inte anpassat efter dina val.
Har du frågor om kursen?
Om du är registrerad på en aktuell kursomgång, se kursrummet i Canvas. Du hittar rätt kursrum under "Kurser" i personliga menyn.
Är du inte registrerad, se Kurs-PM för DD2431 eller kontakta din studentexpedition, studievägledare, eller utbilningskansli.
I Nyhetsflödet hittar du uppdateringar på sidor, schema och inlägg från lärare (när de även behöver nå tidigare registrerade studenter).
Bump. Please do this a couple of days in advance + slides.
In Lecture 1 we will give instructions on it as well as warming-up with Nearest Neighbor classifier (for which no particular reading is required). Welcome to the course!
Will you post the suggested chapters online?
I'm reading on wikipedia right now since neither of the course books have anything on artificial neural networks and there still, unbelievably, aren't any reading instructions available on one page (and this is not to say somewhere inside the schedule)
I think the schedule is pretty readable, once you find it.
Is there some reading for lecture 9, on artificial neural networks? The books don't cover this topic and there is nothing mentioned in the schedule. Or do we have to ask our best friend Google?
@Michiel Van Der Meijden: examination questions don't go beyond the lectures or a literature recommended at corresponding lectures. For curious, there is an online introduction to artificial neural networks by Raul Rojas.
Hey, I wonder if you could publish the Lecture plans? There is no information available.
thank you in advance
Mona
Hi!
Before they upload the info for this years course, you can go to the schedule and choose to view last years lecture plans with slides and readings. Maybe it's not 100% correct but I don't think it has changed dramatically.
@Mona Dadoun: Indeed, we don't publish preliminary plan of the forthcoming lectures but you can always check the last year's schedule (thank you, Michael, for reminding us!). Tip: In order to get the slides, go to the KTH Social of the course and change the view range to the last year.
Thank you both of you!
Hello!
In Lab 1, assignment 3 the code says to run
t.buildTree(m.monk1)
d.check(t, m.monk1test)
But that code will not run as buildTree is missing a parameter (attributes). Shouldn't this be
t.buildTree(m.monk1, m.attributes)
d.check(t, m.monk1test)
?
Hi Casper!
You seem to be right using 't.buildTree' with 'm.attributes'. Thank you for correction!
In the Matlab version you are asked to implement "ent" and "gain" even though they are already implemented in two of the files in the provided directory. In the Python version, you are not asked to implement them but only to use them. What is expected?
Dear Mattias and other participants of this course! Please understand that we cannot answer programming questions in KTH Social on a daily basis. We have reserved Wed 16/9, 17:00-18:00 drop-in hour for help with labs (no booking required). Inquiries regarding Lab 1 welcome to be raised during this time.
Hi!
Atsuto mentioned that it's possible to ask the TAs questions about the labs at some given occasions/time slot. I can't find those time slots... When will they be up un social and/or in schedule?
We will put information about consultations up on Course wiki / Lab booking. The next drop-in hour for help with labs will be 16/9 17:00-18:00, same place as for the lab examinations, Lindstedtsvägen 24, floor 5.
Do you really intend that the only chance to ask questions about lab1 is the day before the bonus deadline? This seems a bit harsh to me, or am I missing something? Is there some other way to get help with the labs?
If the questions are critical for you to complete the lab, maybe you can try to list them here?
I dont get it are there lab exam besides the lab assignment. Or what will be doing in the 10 minutes slot
Lab examination is a 10-minute report of your lab results to a teacher. The teacher decides whether to accept results or not.
Where is it taking place? The location.
When will the final version of Lab 3 be published? Thank you
Lab examinations take place at Lindstedtsvägen 24, Fifth Floor. Meeting at the lunch room. Ring the bell if the door is closed. For details, see page
Lab 3 can be modified during the course term. Go ahead with the old version if you decide to do this exercise earlier.
Hi!
Is it mandatory to do Lab 2 in Python, or are we allowed to use Matlab?
Any programming environment can be used to get results and answer all questions in the lab descriptions. Program manuals are there for your convenience.
Hello Alexander,
I tried to contact Professor Atsuto but failed. My problemis that the first lecture I attended was the one on the 15 because I have arrived to Stockholm late due to the delay in receiving my Residence Permit Card. I am working on catching up with the course material however, I need to extend the assignment and lab examination deadlines considering my circumstances. Please advise.
Thanks
Dear Ahmed,
Relax. Everything will be okay.
Thanks
Hi! Is there a plan to update lab3 to a python version and if so when can we except to find this updated version? :) Thanks!
Lab 3 can be updated at any time. Do it whenever you like, just be consistent and complete the same instructions you started with.
Hi! Regarding what Elizaveta wrote earlier, what we would like to know is if there is going to be a Python version of the instructions and files and when, in that case, those will be ready. Because if there is going to be new instructions we would wait for them, since we would much prefer to do it in Python, but of course we don't want to wait for nothing.
Hi. A Python version is coming very soon.
Hi, it's released now - see above! It is encouraged to check this new version. Matlab version is also available.
Best wishes
Regarding the matlab instructions on lab 3, if you look at the signature for adaboost, isn't the "T" representing the number of classifiers used, and not the number of hypothesis we use (which is 2, since it represents our classes?)? Or am I wrong? Thanks
Hi Johan,
Well "a hypothesis" is a division of your space in two parts; where points in one part are (according to the hypothesis) classified as class 1, and in the other part as class 2. The hypothesis will thus give you one of two possible classifications for each point, but that does not imply there are only two ways of doing this division.
Example of a hypothesis: "All points below the line y = 2x are class 1 and those above it are class 2."
If you have several different divisions, of your input space those are thus different hypotheses.
So yes T is the number of classifiers, but since each classifier will generate one hypothesis (division of your input space), it also equals the number of hypotheses.
All best
Ylva Jansson
I don't understand some of the lab 3 instructions for python. In assignment 1 apparently sigma is supposed to be a three dimensional matrix. However in equation 9 it would appear that each element in sigma is just a scalar, and that sigma is therefore just a 1-dimensional vector. If I check the matlab instructions sigma is instead a two dimensional matrix.
Would be grateful if someone could help me. What am I missing?
I also have a lot of problems with the dimensions of the matrices in Python lab 3 description. If I follow the instructions in the python code of what the matrices whould look like (mu and sigma) iI can't plot gaussians.
Is it possible that equation 8 in lab 3 (python) is wrong? The results seem odd when implemented according to this formula, after replacing N with N_k the plot seems to be as expected.
Yes, this is wrong. It should be N_k for the mean as it is for variance, it has been corrected in a new version. Thank you for spotting it!
Hi Elizaveta and Olof, Equation 9 describes how to estimate the covariance matrix for a single class. It might be confusing here that we use standard vector format (dx1) while the data points might be laid out differently in our data matrix. The result for each class should indeed be a covariance matrix. Then they should be stacked to create a 3-dimensional matrix (or array in numpy) with the given dimensions.
There have been some question from a student regarding the Python version of lab 3.
Q1: In equation 9. Sigma is scalar but in the lab function it is a 3-dim matrix. How does this work?
Answer: This is a typo. Somewhere in the text we mention that x is a row vector, but equations involving x are written as if x was a column vector. For example, in Eq.9 if x is a 1xd vector the covariance estimate should be (x-mu)^T(x-mu) and then the estimate becomes a dxd matrix. The same is true for Eq. 10 and 15. So depending how your internal data structure looks the transpose will be different. So fast solution switch the transpose in the equations if you use x as a row vector. This will also fix the problems for the students having problem plotting the solution.
Q2: I can't get the Olivetti dataset to work properly why is that?
Answer:There is bug in the underlying dimensionality reduction. A variable not being set correctly. This will be fixed in tonight's bug fix.
Q3: Is it possible that equation 8 in lab 3 (python) is wrong? The results seem odd when implemented according to this formula, after replacing N with N_k the plot seems to be as expected.
Answer: Correct. This is fixed in the update that will be out tonight.
We will fix this plus some other bugs in a bug fix that will be out tonight.
Will you release a revised version of "BAYESIAN LEARNING AND BOOSTING"? At the moment there seems to be rather many typos etc.
Will there be a "Debugging help" released for python as well?
Hi,
Two questions about lab 3, assignment 2.2:
1. What is L.H in x = np.linalg.solve(L.H,y) ?
2. Should the discriminent function in equation 10 be a scalar? In that case, is the multiplication in the last step supposed to be a dot product?
Thanks
Answers to questions from Sandra:
1. L.H is "returns the (complex) conjugate transpose of self" for a matrix with real values this is equivalent to L.T. Read about the cholesky decomposition here https://en.wikipedia.org/wiki/Cholesky_decomposition
2. I am not quite sure i understand the question. Eq. 10 is the log posterior. Ask yourself is the log posterior a function that returns a scalar or a vector? (In the old lab version Eq.10 was written as if x was a column vector in the new version the notation is switched to be a row vector. Maybe this is where the confusion arises? )
Hello,
Where can I submit the assignment?
Thanks
Hi, We are having some problems with the Python version of Lab 3.
For example, both equation 12 and 13 seem to lack some information, or contradict themselves.
- The "Ax=b", does that actually refer to the other variables A, x and b used in the factorization to get the inverse covariance? (L = .., y = .., x = ..)
- Is v the resulting inverse covariance?
- What is y?
- What is l_i,i in equation 13?
Seems a bit confusing to us.
Book your time at:
https://www.kth.se/social/course/DD2431/page/lab-booking-2/
If there are no free slots, join someone who presents alone (add your name to the corresponding booking in the table).
Thanks for the answers to my previous questions! I'm still a bit confused about sigma in equation 9 though. According to the equation each element of sigma now corresponds to a d-by-d-matrix. So sigma should be a C,d,d matrix. But what we are supposed to have is a d,d,C matrix. Do we transform by simply transposing or some other method?
Hi Olof,
No sigma should be a d,d,C matrix. We are assuming that sigma is a numpy array. The covariance for the k-th class can then be accessed as
sigma(:,:,k)
Often it is useful to look at the code of the provided files to see how we assume your data structures will look like.
Hi!
What is the assumption when it comes to h (the output in classify())? That is we return a python list and it works in testClassifier(), but not in plotBoundary(). We are doing assignment 2 so its not explicitly mentioned that we should plot boundary, but we thought it would be fun.
Hi, Anton
(1) The instructions says "In Python the code for the generic problem Ax = b" this means that for a general problem when A is factorizable and you are trying to solve the equation system Ax=b then you can write it like that. It is your assignment to try to figure out how to use it in your case.
(2) From ln(|L||L|) it should be clear what l_(i_i) is. If not look in your first year linear algebra book on how to compute the determinant. I am sure there is a case in there where they compute the determinant for a triangular matrix.
Also, when I run testClassifier() on the dataset vowel I get an error message that my sigma and mu are empty. testClassifier() works for iris and wine (I wasn't able to run olivetti on my computer). Vowel is different from all other datasets because its classes don't come in a sequence (all other data comes first with 0-labels, then 1-labels. That is, first 0 0 0 0..., then 1 1 1, etc. Vowel is 0 1 2 3 ... 0 1 2 3) - can you fix that? Or should our mlParams and computePrior be independent of the sequence of labels?
Q0. I have updated the plot boundary function(update will be ready very soon) to better handle cases where the boundary is complicated. However, note that in the case of the olivetti data for example, plotting it in 2-D might not be so helpful since we have taken a 4096 long vector and projected it onto 2 dimensions.
Q1. In this lab you are expected to work with numpy arrays. In the case of the boosting it makes sense to make a list of numpy arrays.
Q2. I cant debug your code. Have you updated to the latest version? There was a debug in the first such that the dimensionality wasn't reduced properly. This is fixed in the latest version.
Q3. As a data scientist you cannot expect the data to be ordered nicely. Hope for the best and plan for the worst!
Tack Martin!
I think there also may be a mistake in labfuns.py. plotGaussian should call plot_cov_ellipse(sigma[:,:,label], mu[label]) and not plot_cov_ellipse(sigma[:,:,0], mu[label]) or am I wrong?
You are correct it should be plot_cov_ellipse(sigma[:,:,label], mu[label]). Fixing this.
Hi,
Just below equation (11), it says that we should solve Σk y = (x − μk )T in order to extract Σ−1. Is x here the "new" values (not the same ones as we used to define Σ and μ). Should it be x* instead? If not, then the function classify() would need another input value.
Thanks.
Hi Frida, (x - mu_k) indeed corresponds to the corresponding expression in the discriminant function. So x^* might have been a clearer notation here, we'll look into changing that.
Hi,
We have a question about lab3, assignment 5.1:
How are we supposed to take the weights into account when computing the priors? Since there are more weight values than prior values.
Thanks
Hi, Sandra. As it says in the lab description, we can look at the weights as taking a particular training point x_i into account w_i times. So if previously there was N_k points in a particular class, we now have to look at "how many times we count" each point. Note that the prior probabilities should still sum to one.
Hello all.
We're trying to solve lab. 3 exercises, but we are still struggling to understand the dimensions for the sigma matrix from eq. 9. As far as we can understand, it should be a C x d x d matrix, and not a d x d x C as is in the description of the exercise. Are we getting something wrong in the way we are interpreting the equation?
Hi Diego,
Eq. 9 is a dxd matrix. In the code you were given we assume that you store the different class sigma matrices as a dxdxC matrix, that is, a dxdxC numpy array.
For the boosting we assume a python list of dxdxC numpy aarrays.
the proposed 3rd step of adaboost needs some adaption in my opinion. During my test runs it occurred that some hypothesis classify "perfectly" and therefore epsilon^t is 0. log(0) -> undef and so is alpha^t. This is propagated to the next iteration and can kill the classify function via undefined values in the covariance matrix.
I solved this by setting alpha^t to +infinity and skipping the remaining iterations. Is this approach appropriate or should I solve that differently?
Hi Stefan,
I think this is a good solution. Clearly, no further improvement can be given by the boosting. And we really appreciate when students take initiative to solve cases of model failure. Another interesting case is to think about when the initial run of the boosting, has a classification accuracy below 50%.
Hi I have a problem in lab 3 for the Matlab code.
I'm running the adaboost functions for the debugger data. The two first iterations is the same compared to the dubberg file. However when I'm getting to the third iteration my, my and sigmas are not the same compared to the debugging pdf file. This feels a bit strange since it works for the two first iterations, anyone having the same problem?
Hi everyone,
I have a problem with the plotBoundary function - the boundaries that it is plotting are extremely complicated and feel incorrect (attached file). This is a sample boundary for iris, with covdiag = False and no boosting. The success rate of the classification under these parameters is 97.5% with a standard deviation of 1.95 over 100 trials and I am getting expected behaviour with respect to boosting and covdiag adjustments for this and other datasets, so I believe that my classification functions is correct.
Is anybody experiencing a similar problem, has any advice, or can plot boundaries that are less complicated?
Thanks, cheers,
Fernando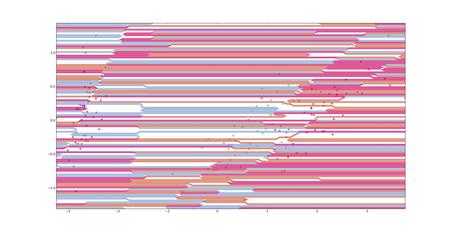
Hi, Fernando
Have you printed how your covariance matrices looks like? Do they look reasonable? Also it is easier to debug with a diagonal covariance matrix if that looks weird then its probably something wrong in your data structure. If it looks ok then there is probably something wrong in your covariance computations.
Hi, Elin
I had a similar problem for a while, check the sum of w_{t+1}^m the at the end of the loop. For me it was coming from a calculus mistake when I was normalizing my w_{t+1}^m.
Hope it helps !
I'm having some problem with the plotBoundary function. It seems like it is sending a matrix with an invalid format on the line:
grid[yi,xi] = classify(np.matrix([[xx, yy]]),prior,mu,sigma,covdiag=covdiag)
in the function plotBoundary.
My classifier seems to be working correctly as I am getting accuracy rates of up to ~ 97%, so for the testClassifier it works correctly.
The error seems to be in the dimensions of X, as the error I am getting is that I am trying to compare the return value of my discriminant function (which should be scalar) with other return values, but it is a vector with two elements when called by plotBoundary rather than a scalar (which it is in all other cases). Does anyone have an idea of why this is occuring and how I can fix it?
I had the same problem. Perhaps you need to adapt your classification-function to accept numpy matrices or convert them before continuing. And furthermore this function does expect a single value as return value instead of any array, matrix ect. So make sure to return a single value if only one value is given via X.
I had the same problem because I use numpy arrays whereas plotBoundary creates numpy matrix so I changed grid[yi,xi] = classify(np.matrix([[xx, yy]]),prior,mu,sigma,covdiag=covdiag) to grid[yi,xi] = classify(np.array([[xx, yy]]),prior,mu,sigma,covdiag=covdiag)
We get the following error when trying the olivetti dataset: "ValueError: zero-size array to reduction operation minimum which has no identity.". Everything works for the iris and vowel datasets. Does anyone know what the error is?
We also have the problem that the wine dataset takes ages to plot (approx. 1 day), but is quick to classify.
Hi Hampus,
Did you update the provided code to the latest version? The first version of the lab had an error were the dimensionality reduction would output a zero-dimension dataset. I think this is what you might be experiencing.
Secondly, the ages to plot-bug have also been fixed. This is was due to not adapting the grid on which we evaluate the classifier on to the particular dataset.
Cheers
M
That fixed it, thanks!
Can someone help me please I'm still struggling with lab 2
I don't understand this part in plotting the Decision Boundary:
What we will have to do is to call your indicator function at a large number of points to see what the classification is at those points. We then draw a contour line at level zero, but also countour lines at 1 and 1 to visualize the margin.”
What large number of points? Should I call the randomized points again to get new values?
And in the indicator(x,y) is x the point of the vector and y the class 1 or -1?
I'm writing this so that anyone experiencing the same problem can find it. At least in my python version returning an N-by-1 matrix from the classify function, according to the instructions, gives the wrong classification result. The reason is that the line which evaluates the performance in the testClassifier function uses yPr==yTe, to check how many predictions are correct. This operation gives a square matrix as a result instead of the desired vector. For me it's working fine by just returning a 1-by-N vector from the classify function instead.
Hope this helps someone!
Hi Elin,
First you need to understand what the decision boundary is representing. You are training a classifier on a set of data points in 2D space, R^2. This classifier will classify points as belonging to either 1 or 0. The data points you are training on are set of samples from the continuous 2D space, R^2. Since you don't know the true distribution of the data you want to evaluate your classifier on a grid in R^2. From this evaluation you can see if the classification is reasonable or not, and detect other qualities of your solution. This classification allows you to draw the decision boundary, that is the boundary that you will have to cross in order to classify a point as 1 instead of 0, or vice versa. Hope this helps.
Hi Olof,
I think there is a bit confusion of using numpy data structures. We are assuming that you are using the standard ndarrays when representing matrices and vectors, and not the numpy matrix structure. This is because the matrix structure is not very flexible with respect to what we are trying to do. In the numpy manual you can read the following on numpy matrices: "A matrix is a specialized 2-D array that retains its 2-D nature through operations." This should explain the error you were getting. We are updating the lab instructions to avoid this confusion for people not familiar with numpy.
Cheers
M
Will there be any more opportunities to present labs? The last time it was fully booked when I wanted to book...
Hello,
My grades from lab 1 and lab 2 still did not appear on rapp. I presented both labs before the bonus deadline. What happened?
@Hugo Sandelius: Join those students who have booked time for one person and present together. If you are going to present lab3, for example, there are 4 time slots like that on Fri 23/10.
@João Marcos Rodrigues: your results are reported now.
@Alexander Kozlov: Okay, but there doesn't seem to be any booking page available at the moment?
It is the same page as before:
https://www.kth.se/social/course/DD2431/page/lab-booking-2-2/
I also added you to the current course list (you weren't there as an old student).
Okay, thanks, I can't access that page though, it says "The page has restricted access. The address given is protected by one of the group's administrators."
It should be readable now. Please make your reservation or write to me directly and I will do this for you.
Hello,
I was checking my results from the labs on rapp, but my grade from lab 2 is still not there. I presented it before the bonus deadline. Could you please tell me when it is going to be updated?
Teachers are filling in Rapp results. We should be ready by the exam week.
How will the lab-bonus be represented in the exam?
Usually bonus points are added to the exam points and the final result is evaluated.
Hi
I registered for the course on Rapp recently. I haven't been graded for lab1 & lab2. I will be presenting lab3 tomorrow. Please look into it
The exam was two weeks ago and we presented lab 3 over three weeks ago. Still nothing in rapp. At this point I'm scared something has been lost in the system. What's going on?
Hi, the results are meant to be reported within 3 weeks time from the date of exam (and it can take the full time given the large number of subscribers). The lab activities will be reflected in the mean time if not appearing yet. Best Wishes.
Oh, to clarify: I'm not talking about the exam results in rapp. I know that takes time. I'm worried about the lack of lab results.
As a comparison, the first 2 labs were reported in less than a day. This one has not been entered even after over three weeks.
Hi Erik, do you remember which of the TA:s you presented to? Was it me or someone else?
Cheers
Martin
I unfortunately don't remember the name of any of the TAs. We presented the lab to a woman.
Hi, then it was me. I do have your name on my list, and it should have been reported ackording to my notes. I will look into what went wrong.
But, your result is not lost, and this will be fixed very soon!
All best
Ylva
Now it has been confirmed and corrected for the following students:
Katelyn Hicks - lab3 15/10 at 14:30
Erik Ihrén - lab3 15/10 at 14:300
Emelie Kullman - lab3 15/10 at 14:10
Gustav Stenbom - lab3 15/10 at 14:10
according to their individual requests.
Hello!
My results for Lab 2 presented Friday 23/10, 09:20 to Nils Bore do not seem be on Rapp yet
Ok, I will have a look at my notes.
@daniel I can confirm that I could not find you in RAPP. That was because I could not find you in the system at the time of the examination. However, I have reported your result to the assistant teacher, so your results are noted with him. No worries.
@Daniel: yes, your lab2 results are registered, now in Rapp too. You took the exam last year and have completed the course by now, as far as I can see.
Hi,
I presented the Lab 3 to Ylva and the result is not on Rapp yet.
@João Marcos Rodrigues: fixed now for you and Diego as well. Thanks for pointing this out!
Hey,
My points for Lab3 are not yet in the RAPP. I was wondering when are they going to be registered?
Best regards, Ana Marija!
@Ana Marija Selak: when and to whom did you present lab3? I can't find your booking time.
Hmm, I am not sure anymore, where can I check that? :/ I was in group with Žad Deljkić and our group name is sinisa vuco. Is that helping?
Yes, full names always help! Fixed now for you two, both in Rapp and in the course report.
Thank you! Best regards
I haven't got any of my lab points yet.
Elin Viberg
@Elin Viberg: you have all lab reports registered now.
Hi!
My points for Lab 3 are not yet in the RAPP. I have presented to TA Ylva Jansson.
@Ezeddin Al Hakim: fixed now.
Lärare Atsuto Maki redigerade 10 september 2015
ExaminatioTentamen
Schemahandläggare redigerade 11 september 2015
ExaminatioTentamen
Schemahandläggare redigerade 11 september 2015
D41, D42, E31, E32, E33, E34, E51, E52, E53, K53, L21, Q22, Q36
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 12, Summary
Schemahandläggare redigerade 31 augusti 2015
[{u'user_name': u'Giampiero Salvi', 'user_id': u'u12rf6rn'}, {'user_name': u'Atsuto Maki', u'user_id': u'u1elx760'}, {'user_name': u'\xd6rjan Ekeberg', 'user_id': u'u1oppomp'}]
Lärare Giampiero Salvi redigerade 8 oktober 2015
Slides for Sequential Data: Lecture 12 Sequential Data handouts¶
Lärare Atsuto Maki redigerade 9 oktober 2015
Slides for Lecture 12¶
Slides for Sequential Data: Lecture 12 Sequential Data handouts
Hello, could you please share also the ConvNet and Reinforcement Learning lecture slides?
Lärare Giampiero Salvi redigerade 12 oktober 2015
Slides for Lecture 12
Slides for Sequential Data: Lecture 12 Sequential Data handouts¶
Lärare Atsuto Maki redigerade 12 oktober 2015
Slides for Lecture 12
Note: The scope of the exam is what has been covered in Lecture 1-11.¶
Hello. I am interested in deepening my knowledge about the topics that were discussed during the summary lecture. Could you please give some reading instructions about those? Thank you.
The last slide of the summary lecture gives a list of related courses taught at KTH. Please check corresponding course pages for the prerequisites and recommended reading. Note also that the material of the last lecture is not included to the exam.
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 8, Classification with Separating Hyperplanes
Schemahandläggare redigerade 31 augusti 2015
[{u'user_name': u'Giampiero Salvi\xd6rjan Ekeberg', u'user_id': u'u12rf6rnoppomp'}]
Hi! Could you please share the slides from your lecture and also which chapters from the course literature should we read?
@Elizaveta I think that you should be reading Chapter 9 for this lecture here.
Lärare Örjan Ekeberg redigerade 28 september 2015
Slides from lecture 8¶
Lärare Örjan Ekeberg redigerade 28 september 2015
Topics:¶
* Linear separation in high dimensional spaces
* Structural risk minimization
* Support Vector Machines
* Kernels for separating in a higher dimensional space
* Non-separable classes
Slides from lecture 8
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 6, Probability I
Lärare Atsuto Maki redigerade 28 augusti 2015
Lecture 6, Probability II
Schemahandläggare redigerade 11 september 2015
MF2
Lärare Giampiero Salvi redigerade 17 september 2015
Recommended reading:¶
Prince, Chapter 3, 4, 7.4. (Book available in full PDF here)¶
Lärare Giampiero Salvi redigerade 17 september 2015
Topics:¶
* estimation theory
* Maximum Likelihood, Maximum a Posteriori, Bayesian estimation
* Unsupervised learning and K-means
* Mixture of distributions and Expectation Maximization algorithm
Slides for Lecture 06¶
Recommended reading:
Prince, Chapter 3, 4, 7.4. (Book available in full PDF here)
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 5, Probability I
Schemahandläggare redigerade 31 augusti 2015
[{u'user_name': u'Atsuto MakGiampiero Salvi', u'user_id': u'u1elx7602rf6rn'}]
Lärare Giampiero Salvi redigerade 15 september 2015
Topics:¶
* concepts of probability theory
* random variables and distributions
* conditional probabilities and Bayes rule
* Bayesian inference
Slides for Lecture 5¶
Related reading:¶
Prince, S.J.D., Part I (Chapters 2, 3, 5)¶
If you want to know more a great book on these topics isBishop, C. M. Pattern Recognition and Machine Learning,Springer. ¶
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 2, Decision Trees
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
¶
Related reading:¶
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)¶
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/¶
Lärare Atsuto Maki redigerade 28 augusti 2015
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
¶ Related reading:
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 28 augusti 2015
Topics:¶
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
Related reading:
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Schemahandläggare redigerade 1 september 2015
Torsdag 3 september 2015 kl 137:00 - 159:00
B3E1
Flyttad från kl 13-15 pga för liten sal
Lärare Atsuto Maki redigerade 2 september 2015
Topics:
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
Slides for Lecture 2¶
Related reading:
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 3 september 2015
Topics:
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
Slides for Lecture 2
Related reading:
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 27 oktober 2015
Topics:
* What is a Decision Tree?
* When are decision trees useful?
* How can one select what questions to ask?
* What do we mean by Entropy for a data set?
* What do we mean by the Information Gain of a question?
* What is the problem of overfitting? Minimizing training error?
* What extensions will be possible for improvement?
Slides for Lecture 2
Related reading:
Chapter 8.1 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 10, Ensemble Methods
Schemahandläggare redigerade 31 augusti 2015
KM1
Lärare Atsuto Maki redigerade 29 september 2015
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10¶
Related reading:¶
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)¶
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/¶
Lärare Atsuto Maki redigerade 2 oktober 2015
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10
Slides for Lecture 10 (full size)¶
Related reading:
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 2 oktober 2015
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10
Slides for Lecture 10 (full size)
Related reading:
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 5 oktober 2015
Topics:¶
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10
Slides for Lecture 10 (full size)
Related reading:
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 6 oktober 2015
Topics:
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10
Slides for Lecture 10 (full size)
Related reading:
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 26 oktober 2015
Topics:
* Why combine classifiers?
* Bagging
* Decision Forests
* Boosting
Slides for Lecture 10
Slides for Lecture 10 (full size)
Related reading:
Chapter 8.2 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 28 augusti 2015
FöreläsningLecture 7, Classification Introduction
Schemahandläggare redigerade 31 augusti 2015
[{u'user_name': u'\xd6rjan EkebergAtsuto Maki', u'user_id': u'u1oppompelx760'}]
Lärare Atsuto Maki redigerade 17 september 2015
Related reading:¶
Chapter 4 from An Introduction to Statistical Learning (Springer, 2013)¶
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/¶
Lärare Atsuto Maki redigerade 17 september 2015
Topics:¶
* Naive Bayes
* Logistic regression
* Inference and decision
* Discriminative function
* Discriminative vs Generative model
Related reading:
Chapter 4 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 21 september 2015
Topics:
* Naive Bayes
* Logistic regression
* Inference and decision
* Discriminative function
* Discriminative vs Generative model
Slides for Lecture 7¶
Related reading:
Chapter 4 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Lärare Atsuto Maki redigerade 21 september 2015
Topics:
* Naive Bayes
* Logistic regression
* Inference and decision
* Discriminative function
* Discriminative vs Generative model
Slides for Lecture 7
Slides for Lecture 7 (part II)¶
Related reading:
Chapter 4 from An Introduction to Statistical Learning (Springer, 2013)
Gareth James, Daniela Witten, Trevor Hastie and Robert TibshiraniAvailable online: http://www-bcf.usc.edu/~gareth/ISL/
Will you be posting reading assignments (i.e. what to read prior to each lecture etc)?