Experimental Genomics
The research focuses on development of molecular methods allowing large-scale analysis of DNA, RNA and proteins in both bulk samples and single cells.
Research
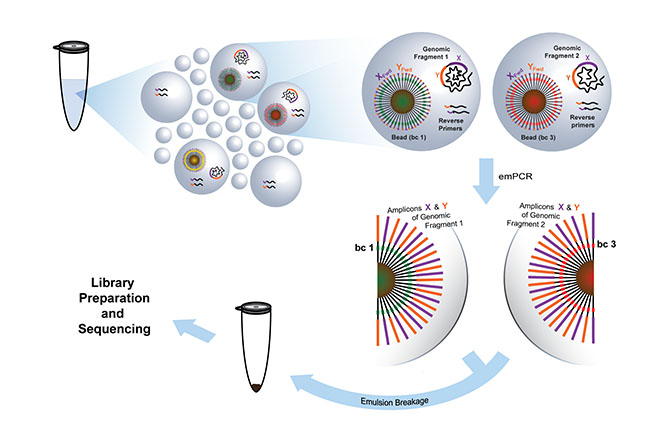
A method that has facilitated high throughput investigation of the functional constituents of a cell is massively DNA barcoding of emulsion droplets. In this method, the DNA barcodes are linked to DNA (or cDNA) molecules in droplets and sequenced using a massively parallel sequencing (MPS) platform.
The sequenced DNA molecules are each traced back to a share droplet allowing mutation analysis and transcript profiling of individual cells. The group is also working on a new approach for droplet-based DNA-assisted proteomics. The main focus of the project is on a novel principle for simultaneous detection and quantification of proteins in complex samples, where a combination of immunorecognition (DNA-labeled antibodies) and massively parallel DNA sequencing is applied.
The research group has previously developed trinucleotide threading (TnT), protease-mediated allele-specific extension (PrASE) and apyrase-mediated allele-specific extension (AMASE) to assess DNA variations and to profile expressed gene in a multiplex fashion.
