Protein-based prodrugs
Proteolytic processing is a common means in nature for regulating activity of proteins. We are investigating strategies for controlling activity of engineered affinity proteins by a similar mechanism.
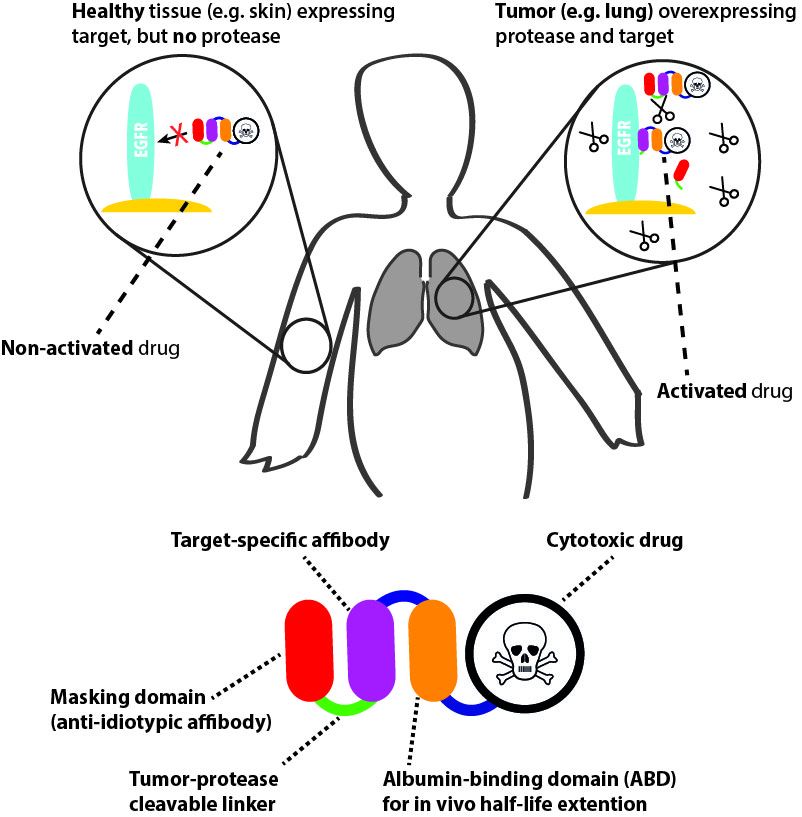
The project is based on a dimeric form of “self-blocking affinity proteins”, in which the first domain is specific for a target, while the second binds the first domain and blocks its interaction with the target. The domains are fused by a linker with a protease site, and target binding can be regulated using proteolytic digestion. The concept might be exploited for several purposes (e.g. biosensors etc.). An interesting field is tumor targeting and we believe we have potential to improve targeting agents to improve tumor selectivity. Many tumors show increased levels of proteases, which might be exploited for activation of prodrugs. The hypothesis is that the prodrug will be inert (blocked) in normal tissues, while the blocking domain will be cleaved off in the tumor, hence sparing normal tissues and decreasing side effects. Long term, this will potentially result in improved targeting agents, allowing for reduced toxicity and higher therapeutic effect and the possibility of targeting tumor antigens that are currently “non-druggable” due to toxicity.
Research funded by: Vinnova - Sweden's innovation agency and the Swedish Research Council