Christofer Lendel
Professor
Details
Researcher
About me
Christofer Lendel received his PhD in Biotechnology from KTH in 2006. After post-doctoral research at the University of Cambridge, Stockholm University and the Swedish University for Agricultural Sciences in Uppsala he returned to KTH in 2015. He has currently a position as Professor in Biophysical Chemistry with a focus on materials and interfaces.
Christofer Lendel’s research concerns the structural and functional properties of protein molecules; i.e. what they look like, how they perform their functions, why they sometimes misbehave and cases disease, and how we can exploit them as building blocks for biobased materials. In particular, he is fascinated by the self-assembly of protein molecules into nanoscale fibrils. These are key events in many biochemical processes ranging from neurodegenerative disorders, such as Alzheimer’s disease, to the design of novel nanomaterials. One of the central research questions is whether there are any fundamental differences between the fibrils produced from agricultural protein resources and those that form in our bodies when we suffer from disease. Another is how we can mimic natural assembly processes for proteins materials in order to create new solutions for a sustainable society. This could be materials that replace environmentally problematic materials (e.g. oil-based plastics) or that have a function that can solve environmental challenges (e.g. water purification). Another important area is the food sector, where the structural properties of proteins constitute a significant part of the taste experience.
Latest news
New insights about the structure of spider silk fibers
In collaboration with My Hedhammar's group we have explored the structurucral properties of spider silk. Spider silk proteins, spidroins, are complex macromolecules consisting of a repetitive region flanked by non-repetitive N- and C-terminal domains (NT and CT). While the repetitive region is fundamental for the mechanical properties of the final fiber, the terminal domains are believed to control the assembly process, as a response on changes in the environment when the spidroins are pushed through the spinning duct. We exploit the ability of the CT domain to form silk-like fibers on its own, to gain new insights about its functional role. Using an integrative structural biology approach, we show for the first time that the CT domain, unlike the NT domain, undergoes an alpha-to-beta structural transition upon fiber formation. The results highlight a segment that is part of helix 4 in the soluble structure of CT as a potential driver for this process. The paper was just published in Nature Communications.
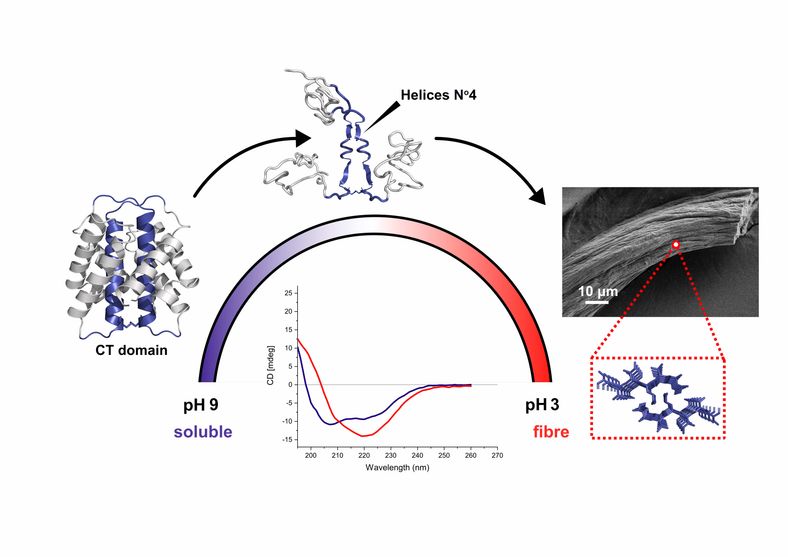
Research Interests
The assembly of protein molecules into nanoscale aggregates and amyloid fibrils are key events in many biochemical processes ranging from neurodegenerative disorders, such as Alzheimer’s disease, to the design of novel nanomaterials. In my research I investigate the structural properties and interactions of such protein aggregates. In particular, I am interested in the molecular mechanisms for how protein nanostructures interact with other (bio)molecules.
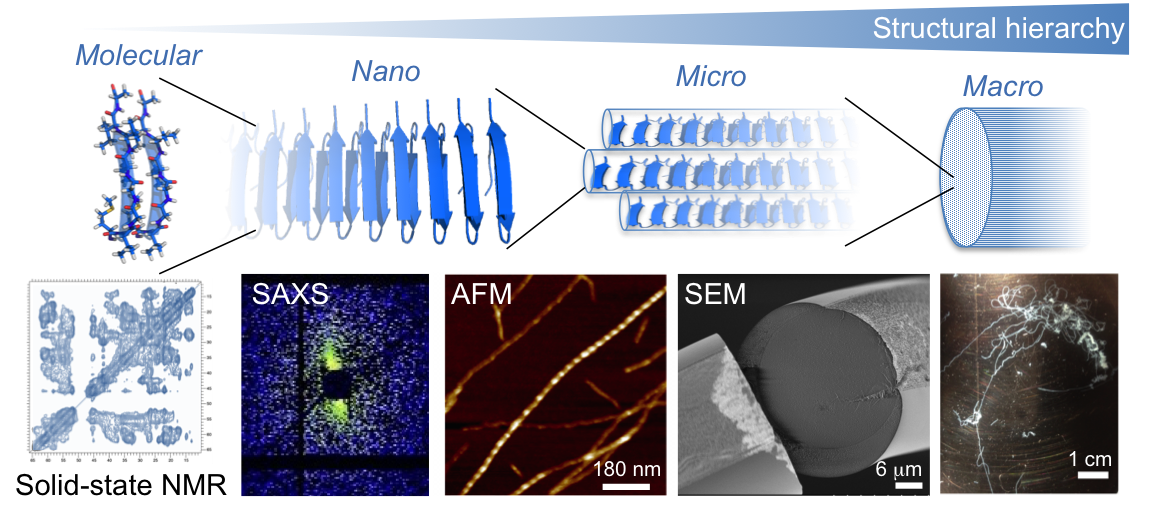
New materials from protein nanofibrils
Protein nanofibrils display extraordinary mechanical and functional properties and has the potential to be used as building blocks for new materials with hieracrical structures. We develop methods to produce fibrils from protein-rich and renewable sources (e.g. plant proteins) and the technology to produce new protein-based materials.
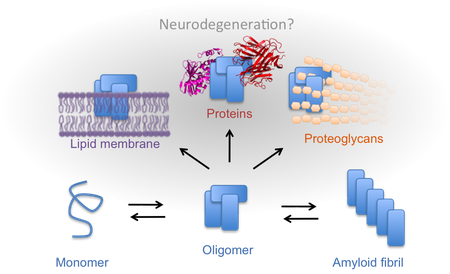
Pathological mechanisms of protein aggregates
Protein self-assembly and deposition are hallmarks of many serious diseases but the disease-causing mechanisms remain enigmatic. However, any pathological mechanism must involve interactions between the protein aggregates and other biomolecules. We explore and characterize such interactions in order to better understand what makes a protein nanostructure toxic.
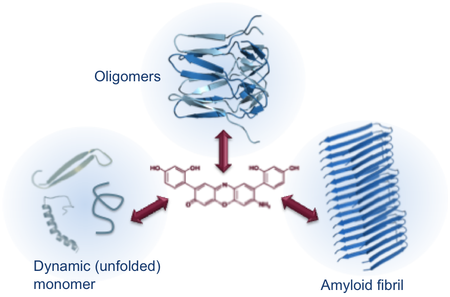
Inhibitors of protein aggregation
The mechanisms of action for small molecule inhibitors of protein aggregation remain poorly understood and the development of new therapeutics against amyloid diseases is slow. Interfering with protein self-assembly is complicated and essentially different from traditional drug design. By studying the binding mechanisms of small molecules to aggregation-prone proteins we believe that we will better understand what type of molecules could be explored as drug candidates for amyloid-diseases.
Group members
Rahul Sharma (Post Doc)
Jing Hu (Post Doc)
Selected publications
Binding of human proteins to amyloid-β protofibrils. Rahmanet al. ACS Chem. Biol. (2015).
Inhibition of amyloid formation (review). Härd & Lendel J. Mol. Biol. (2012).
Courses
Analysis of Biomolecules (BB1200), teacher
Degree Project in Chemistry, Second Cycle (KD200X), examiner
Food chemistry and technology (CK2000), course responsible
Introductory Chemistry (KD1020), teacher
Perspectives on Research and Innovation (KA1030), teacher
Perspectives on Research and Innovation (KA1040), teacher
Project in Chemistry (KD2920), examiner
Project in Chemistry (KD2905), examiner
Research Frontiers in Chemistry (FCK3315), teacher
Thermodynamics (KE1160), course responsible