School of Electrical Engineering and Computer Science
The School of Electrical Engineering and Computer Science (EECS) is one of five schools at KTH Royal Institute of Technology. We conduct research and education in electrical engineering, computer science, intelligent systems and human centered technology. Here you can read about our research, education and the latest news at EECS. Located at KTH Kista and KTH Campus in Stockholm.
EECS' education programmes
The School of Electrical Engineering and Computer Science educates engineers, master students and researchers in electrical engineering and computer science. We are located at two different campuses in Stockholm.
Studies in Electrical Engineering and Computer Science
Doctoral (Ph.D) programmes at EECS
EECS' departments
Latest news from EECS
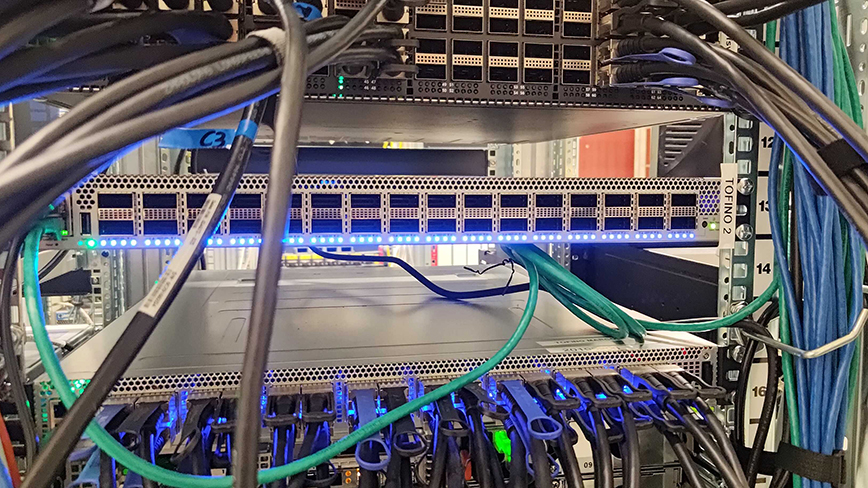

New cybersecurity analysis solution leads to significant reduction in energy consumption
By offloading calculations for complex cybersecurity analyses to network accelerators, energy consumption can be reduced by over 30 times, according to Mariano Scazzariello, a postdoctoral researcher ...
Read the article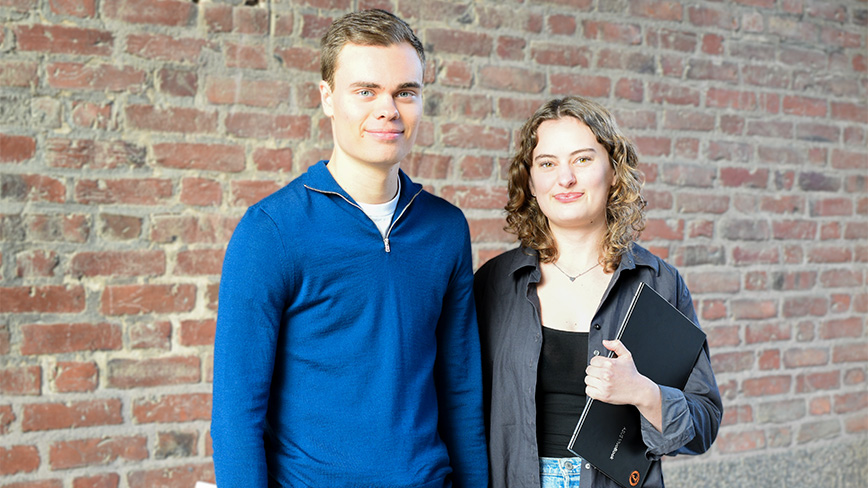
How to stop cyber-attacks with honeypots
In the ever-evolving landscape of cyber warfare, defending against human-controlled cyberattacks requires innovative strategies. A recent study conducted by students at KTH delves into the realm of cy...
Read the article
AI in coding awarded for impact
For a long time, coding was tediously manual, but in 2009, Martin Monperrus, Professor of Software Engineering at KTH Royal Institute of Technology, and his team realised that built-in AI could help m...
Read the articleMore news
- New cybersecurity analysis solution leads to significant reduction in energy consumption
19 Feb 2024
- How to stop cyber-attacks with honeypots
13 Feb 2024
- AI in coding awarded for impact
5 Feb 2024
- Emil is KTH's first cybersecurity engineer
19 Dec 2023
- Research wants to change fetal monitoring at birth
18 Dec 2023
Calendar
-
General
Tue 2024-04-23, 12:00 - Thu 2024-05-02, 12:00
Participating: Anne-Marie Dehon, Bogil Lee, Daniel R Obregón, Davide Ronco, Erik Lindeborg, Henny L Kjellberg
Location: Biblioteket, Konstfack, LM Ericssons väg 14, Stockholm
2024-04-23T12:00:00.000+02:00 2024-05-02T12:00:00.000+02:00 in the making: the crack that lets the light shine in (General) Biblioteket, Konstfack, LM Ericssons väg 14, Stockholm (KTH, Stockholm, Sweden)in the making: the crack that lets the light shine in (General) -
Seminars and lectures
Friday 2024-04-26, 15:00 - 16:00
Participating: Nicolas Tonello, ConstelCom and Renuda, UK
Location: Online via Zoom
Video link: https://kth-se.zoom.us/meeting/61393347130
2024-04-26T15:00:00.000+02:00 2024-04-26T16:00:00.000+02:00 Webinar: Introduction to Code-Saturne (Seminars and lectures) Online via Zoom (KTH, Stockholm, Sweden)Webinar: Introduction to Code-Saturne (Seminars and lectures) -
Public defences of doctoral theses
Monday 2024-04-29, 13:00
Location: Kollegiesalen, Brinellvägen 8, Stockholm
Video link: https://kth-se.zoom.us/j/68660447128
Doctoral student: Federico Favero , Medieteknik och interaktionsdesign, MID
2024-04-29T13:00:00.000+02:00 2024-04-29T13:00:00.000+02:00 Light Rhythms (Public defences of doctoral theses) Kollegiesalen, Brinellvägen 8, Stockholm (KTH, Stockholm, Sweden)Light Rhythms (Public defences of doctoral theses)









